Organelle interactions compartmentalize hepatic fatty acid trafficking and metabolism
- PMID: 37104088
- PMCID: PMC10278152
- DOI: 10.1016/j.celrep.2023.112435
Organelle interactions compartmentalize hepatic fatty acid trafficking and metabolism
Abstract
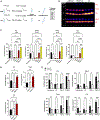
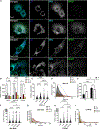
Organelle interactions play a significant role in compartmentalizing metabolism and signaling. Lipid droplets (LDs) interact with numerous organelles, including mitochondria, which is largely assumed to facilitate lipid transfer and catabolism. However, quantitative proteomics of hepatic peridroplet mitochondria (PDM) and cytosolic mitochondria (CM) reveals that CM are enriched in proteins comprising various oxidative metabolism pathways, whereas PDM are enriched in proteins involved in lipid anabolism. Isotope tracing and super-resolution imaging confirms that fatty acids (FAs) are selectively trafficked to and oxidized in CM during fasting. In contrast, PDM facilitate FA esterification and LD expansion in nutrient-replete medium. Additionally, mitochondrion-associated membranes (MAM) around PDM and CM differ in their proteomes and ability to support distinct lipid metabolic pathways. We conclude that CM and CM-MAM support lipid catabolic pathways, whereas PDM and PDM-MAM allow hepatocytes to efficiently store excess lipids in LDs to prevent lipotoxicity.
Keywords: CP: Metabolism; MAM; cytosolic mitochondria; fatty acids; lipid anabolism; lipid catabolism; lipid droplets; organelle interactions; peridroplet mitochondria; perilipin 5; single-molecule localization microscopy.
Copyright © 2023 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures





Similar articles
-
Mitofusin-2 induced by exercise modifies lipid droplet-mitochondria communication, promoting fatty acid oxidation in male mice with NAFLD.Metabolism. 2024 Mar;152:155765. doi: 10.1016/j.metabol.2023.155765. Epub 2023 Dec 23. Metabolism. 2024. PMID: 38142958
-
Peridroplet mitochondria are associated with the severity of MASLD and the prevention of MASLD by diethyldithiocarbamate.J Lipid Res. 2024 Aug;65(8):100590. doi: 10.1016/j.jlr.2024.100590. Epub 2024 Jul 7. J Lipid Res. 2024. PMID: 38981572 Free PMC article.
-
Fasting and refeeding induces changes in the mouse hepatic lipid droplet proteome.J Proteomics. 2018 Jun 15;181:213-224. doi: 10.1016/j.jprot.2018.04.024. Epub 2018 Apr 23. J Proteomics. 2018. PMID: 29698803
-
Mitochondria Bound to Lipid Droplets: Where Mitochondrial Dynamics Regulate Lipid Storage and Utilization.Cell Metab. 2019 Apr 2;29(4):827-835. doi: 10.1016/j.cmet.2019.02.011. Epub 2019 Mar 21. Cell Metab. 2019. PMID: 30905670 Free PMC article. Review.
-
Fatty Acid Trafficking Between Lipid Droplets and Mitochondria: An Emerging Perspective.Int J Biol Sci. 2025 Feb 10;21(5):1863-1873. doi: 10.7150/ijbs.105361. eCollection 2025. Int J Biol Sci. 2025. PMID: 40083687 Free PMC article. Review.
Cited by
-
Integrated systems biology identifies disruptions in mitochondrial function and metabolism as key contributors to heart failure with preserved ejection fraction (HFpEF).bioRxiv [Preprint]. 2024 Oct 25:2024.10.25.619450. doi: 10.1101/2024.10.25.619450. bioRxiv. 2024. Update in: JACC Basic Transl Sci. 2025 Aug 15;10(9):101334. doi: 10.1016/j.jacbts.2025.101334. PMID: 39484400 Free PMC article. Updated. Preprint.
-
Imaging and proteomics toolkits for studying organelle contact sites.Front Cell Dev Biol. 2024 Sep 24;12:1466915. doi: 10.3389/fcell.2024.1466915. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 39381373 Free PMC article. Review.
-
A fluorogenic complementation tool kit for interrogating lipid droplet-organelle interaction.bioRxiv [Preprint]. 2023 Nov 29:2023.11.29.569289. doi: 10.1101/2023.11.29.569289. bioRxiv. 2023. Update in: J Cell Biol. 2024 Sep 2;223(9):e202311126. doi: 10.1083/jcb.202311126. PMID: 38076863 Free PMC article. Updated. Preprint.
-
Diallyl disulfide alleviates hepatic steatosis by the conservative mechanism from fish to tetrapod: Augment Mfn2/Atgl-Mediated lipid droplet-mitochondria coupling.Redox Biol. 2024 Nov;77:103395. doi: 10.1016/j.redox.2024.103395. Epub 2024 Oct 15. Redox Biol. 2024. PMID: 39447254 Free PMC article.
-
Functional compartmentalization of hepatic mitochondrial subpopulations during MASH progression.Commun Biol. 2025 Feb 18;8(1):258. doi: 10.1038/s42003-025-07713-9. Commun Biol. 2025. PMID: 39966593 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

