A genomic timescale for placental mammal evolution
- PMID: 37104581
- PMCID: PMC10233747
- DOI: 10.1126/science.abl8189
A genomic timescale for placental mammal evolution
Abstract
The precise pattern and timing of speciation events that gave rise to all living placental mammals remain controversial. We provide a comprehensive phylogenetic analysis of genetic variation across an alignment of 241 placental mammal genome assemblies, addressing prior concerns regarding limited genomic sampling across species. We compared neutral genome-wide phylogenomic signals using concatenation and coalescent-based approaches, interrogated phylogenetic variation across chromosomes, and analyzed extensive catalogs of structural variants. Interordinal relationships exhibit relatively low rates of phylogenomic conflict across diverse datasets and analytical methods. Conversely, X-chromosome versus autosome conflicts characterize multiple independent clades that radiated during the Cenozoic. Genomic time trees reveal an accumulation of cladogenic events before and immediately after the Cretaceous-Paleogene (K-Pg) boundary, implying important roles for Cretaceous continental vicariance and the K-Pg extinction in the placental radiation.
Conflict of interest statement
Figures

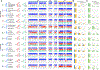



References
-
- Eisenberg JF, The Mammalian Radiations: An Analysis of Trends in Evolution, Adaptation and Behaviour (Univ. Chicago Press, 1981).
-
- O’Brien SJ, Graphodatsky AS, Perelman PL, Atlas of Mammalian Chromosomes (Wiley Blackwell, 2020).

