The Yeast Permease Agp2 Senses Cycloheximide and Undergoes Degradation That Requires the Small Protein Brp1-Cellular Fate of Agp2 in Response to Cycloheximide
- PMID: 37108141
- PMCID: PMC10138708
- DOI: 10.3390/ijms24086975
The Yeast Permease Agp2 Senses Cycloheximide and Undergoes Degradation That Requires the Small Protein Brp1-Cellular Fate of Agp2 in Response to Cycloheximide
Abstract
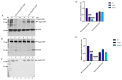
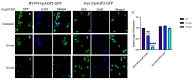
The Saccharomyces cerevisiae Agp2 is a plasma membrane protein initially reported to be an uptake transporter for L-carnitine. Agp2 was later rediscovered, together with three additional proteins, Sky1, Ptk2, and Brp1, to be involved in the uptake of the polyamine analogue bleomycin-A5, an anticancer drug. Mutants lacking either Agp2, Sky1, Ptk2, or Brp1 are extremely resistant to polyamines and bleomycin-A5, suggesting that these four proteins act in the same transport pathway. We previously demonstrated that pretreating cells with the protein synthesis inhibitor cycloheximide (CHX) blocked the uptake of fluorescently labelled bleomycin (F-BLM), raising the possibility that CHX could either compete for F-BLM uptake or alter the transport function of Agp2. Herein, we showed that the agp2Δ mutant displayed striking resistance to CHX as compared to the parent, suggesting that Agp2 is required to mediate the physiological effect of CHX. We examined the fate of Agp2 as a GFP tag protein in response to CHX and observed that the drug triggered the disappearance of Agp2 in a concentration- and time-dependent manner. Immunoprecipitation analysis revealed that Agp2-GFP exists in higher molecular weight forms that were ubiquitinylated, which rapidly disappeared within 10 min of treatment with CHX. CHX did not trigger any significant loss of Agp2-GFP in the absence of the Brp1 protein; however, the role of Brp1 in this process remains elusive. We propose that Agp2 is degraded upon sensing CHX to downregulate further uptake of the drug and discuss the potential function of Brp1 in the degradation process.
Keywords: a small protein; cycloheximide; drugs; protein degradation; protein synthesis inhibitor; sensor; ubiquitinylation; uptake transporter; yeast.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Loss of the yeast transporter Agp2 upregulates the pleiotropic drug-resistant pump Pdr5 and confers resistance to the protein synthesis inhibitor cycloheximide.PLoS One. 2024 May 22;19(5):e0303747. doi: 10.1371/journal.pone.0303747. eCollection 2024. PLoS One. 2024. PMID: 38776347 Free PMC article.
-
Agp2, a member of the yeast amino acid permease family, positively regulates polyamine transport at the transcriptional level.PLoS One. 2013 Jun 3;8(6):e65717. doi: 10.1371/journal.pone.0065717. Print 2013. PLoS One. 2013. PMID: 23755272 Free PMC article.
-
Characterization of a transport and detoxification pathway for the antitumour drug bleomycin in Saccharomyces cerevisiae.Biochem J. 2004 Nov 15;384(Pt 1):47-58. doi: 10.1042/BJ20040392. Biochem J. 2004. PMID: 15248838 Free PMC article.
-
A new twist in cellular resistance to the anticancer drug bleomycin-A5.Curr Drug Metab. 2010 Sep;11(7):595-602. doi: 10.2174/138920010792927307. Curr Drug Metab. 2010. PMID: 20812903 Review.
-
Sky1: at the intersection of prion-like proteins and stress granule regulation.Curr Genet. 2020 Jun;66(3):463-468. doi: 10.1007/s00294-019-01044-z. Epub 2019 Nov 19. Curr Genet. 2020. PMID: 31745569 Free PMC article. Review.
Cited by
-
Mft1, identified from a genome-wide screen of the yeast haploid mutants, mediates cell cycle arrest to counteract quinoxaline-induced toxicity.Front Genet. 2024 Jan 12;14:1296383. doi: 10.3389/fgene.2023.1296383. eCollection 2023. Front Genet. 2024. PMID: 38283148 Free PMC article.
-
Loss of the yeast transporter Agp2 upregulates the pleiotropic drug-resistant pump Pdr5 and confers resistance to the protein synthesis inhibitor cycloheximide.PLoS One. 2024 May 22;19(5):e0303747. doi: 10.1371/journal.pone.0303747. eCollection 2024. PLoS One. 2024. PMID: 38776347 Free PMC article.
References
-
- van Roermund C.W., Hettema E.H., van den Berg M., Tabak H.F., Wanders R.J. Molecular characterization of carnitine-dependent transport of acetyl-CoA from peroxisomes to mitochondria in Saccharomyces cerevisiae and identification of a plasma membrane carnitine transporter, Agp2p. Embo J. 1999;18:5843–5852. doi: 10.1093/emboj/18.21.5843. - DOI - PMC - PubMed
-
- Iraqui I., Vissers S., Bernard F., de Craene J.O., Boles E., Urrestarazu A., Andre B. Amino acid signaling in Saccharomyces cerevisiae: A permease-like sensor of external amino acids and F-Box protein Grr1p are required for transcriptional induction of the AGP1 gene, which encodes a broad-specificity amino acid permease. Mol. Cell. Biol. 1999;19:989–1001. doi: 10.1128/MCB.19.2.989. - DOI - PMC - PubMed
-
- Lee J., Lee B., Shin N., Kwak S.-S., Bahk J.D., Lim C.O., Yun D.-J. Carnitine uptake by AGP2 in yeast Saccharomyces cerevisiae is dependent on Hog1 MAP kinase pathway. Mol. Cells. 2002;13:407–412. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

