Interaction Network Construction and Functional Analysis of the Plasma Membrane H+-ATPase in Bangia fuscopurpurea (Rhodophyta)
- PMID: 37108805
- PMCID: PMC10142769
- DOI: 10.3390/ijms24087644
Interaction Network Construction and Functional Analysis of the Plasma Membrane H+-ATPase in Bangia fuscopurpurea (Rhodophyta)
Abstract
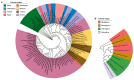
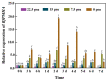
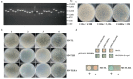
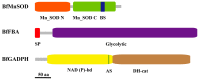
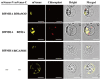
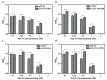
Salinity is a serious threat to most land plants. Although seaweeds adapt to salty environments, intertidal species experience wide fluctuations in external salinities, including hyper- and hypo-saline stress. Bangia fuscopurpurea is an economic intertidal seaweed with a strong tolerance to hypo-salinity. Until now, the salt stress tolerance mechanism has remained elusive. Our previous study showed that the expression of B. fuscopurpurea plasma membrane H+-ATPase (BfPMHA) genes were the most upregulated under hypo-salinity. In this study, we obtained the complete sequence of BfPMHA, traced the relative expression of this BfPMHA gene in B. fuscopurpurea under hypo-salinity, and analyzed the protein structure and properties based on the gene's sequence. The result showed that the expression of BfPMHA in B. fuscopurpurea increased significantly with varying hypo-salinity treatments, and the higher the degree of low salinity stress, the higher the expression level. This BfPMHA had typical PMHA structures with a Cation-N domain, an E1-E2 ATPase domain, a Hydrolase domain, and seven transmembrane domains. In addition, through the membrane system yeast two-hybrid library, three candidate proteins interacting with BfPMHA during hypo-saline stress were screened, fructose-bisphosphate aldolase (BfFBA), glyceraldehyde 3-phosphate dehydrogenase (NADP+) (phosphorylating) (BfGAPDH), and manganese superoxide dismutase (BfMnSOD). The three candidates and BfPMHA genes were successfully transferred and overexpressed in a BY4741 yeast strain. All of them significantly enhanced the yeast tolerance to NaCl stress, verifying the function of BfPMHA in salt stress response. This is the first study to report the structure and topological features of PMHA in B. fuscopurpurea and its candidate interaction proteins in response to salt stress.
Keywords: hypo-salinity; interaction proteins; plasma membrane H+-ATPase; salt stress; seaweed; yeast two-hybrid.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Integrative analyses of transcriptomes and metabolomes provide insight into salinity adaption in Bangia (Rhodaphyta).Int J Biol Macromol. 2023 Dec 31;253(Pt 8):127466. doi: 10.1016/j.ijbiomac.2023.127466. Epub 2023 Oct 22. Int J Biol Macromol. 2023. PMID: 37875187
-
Integrating transcriptomics and metabolomics to characterize the regulation of EPA biosynthesis in response to cold stress in seaweed Bangia fuscopurpurea.PLoS One. 2017 Dec 14;12(12):e0186986. doi: 10.1371/journal.pone.0186986. eCollection 2017. PLoS One. 2017. PMID: 29240755 Free PMC article.
-
Plasma membrane-localized H+-ATPase OsAHA3 functions in saline-alkaline stress tolerance in rice.Plant Cell Rep. 2023 Dec 22;43(1):9. doi: 10.1007/s00299-023-03103-9. Plant Cell Rep. 2023. PMID: 38133824
-
Root-Apex Proton Fluxes at the Centre of Soil-Stress Acclimation.Trends Plant Sci. 2020 Aug;25(8):794-804. doi: 10.1016/j.tplants.2020.03.002. Epub 2020 Mar 30. Trends Plant Sci. 2020. PMID: 32673580 Review.
-
The cellular biology of proton-motive force generation by V-ATPases.J Exp Biol. 2000 Jan;203(Pt 1):89-95. doi: 10.1242/jeb.203.1.89. J Exp Biol. 2000. PMID: 10600677 Review.
Cited by
-
Transcriptome Reveals the Differential Regulation of Sugar Metabolism to Saline-Alkali Stress in Different Resistant Oats.Genes (Basel). 2025 Jan 20;16(1):105. doi: 10.3390/genes16010105. Genes (Basel). 2025. PMID: 39858652 Free PMC article.
References
-
- Butcher K., Wick A.F., DeSutter T., Chatterjee A., Harmon J. Soil salinity: A threat to global food security. Agron. J. 2016;108:2189–2200. doi: 10.2134/agronj2016.06.0368. - DOI
-
- Karsten U. Seaweed Biology. Springer; Berlin/Heidelberg, Germany: 2012. Seaweed acclimation to salinity and desiccation stress; pp. 87–107.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

