Genome-Wide Comparative Analysis of the R2R3-MYB Gene Family in Six Ipomoea Species and the Identification of Anthocyanin-Related Members in Sweet Potatoes
- PMID: 37111954
- PMCID: PMC10140993
- DOI: 10.3390/plants12081731
Genome-Wide Comparative Analysis of the R2R3-MYB Gene Family in Six Ipomoea Species and the Identification of Anthocyanin-Related Members in Sweet Potatoes
Abstract
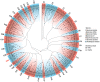
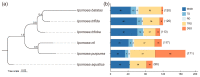
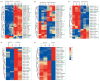
Sweet potatoes (Ipomoea batatas) are one of the important tuberous root crops cultivated worldwide, and thier storage roots are rich in antioxidants, such as anthocyanins. R2R3-MYB is a large gene family involved in various biological processes, including anthocyanin biosynthesis. However, few reports about the R2R3-MYB gene family of sweet potatoes have been released to date. In the present study, a total of 695 typical R2R3-MYB genes were identified in six Ipomoea species, including 131 R2R3-MYB genes in sweet potatoes. A maximum likelihood phylogenetic analysis divided these genes into 36 clades, referring to the classification of 126 R2R3-MYB proteins of Arabidopsis. Clade C25(S12) has no members in six Ipomoea species, whereas four clades (i.e., clade C21, C26, C30, and C36), including 102 members, had no members in Arabidopsis, and they were identified as Ipomoea-specific clades. The identified R2R3-MYB genes were unevenly distributed on all chromosomes in six Ipomoea species genomes, and the collinearity analysis among hexaploid I. batatas and another five diploid Ipomoea species suggested that the sweet potato genome might have undergone a larger chromosome rearrangement during the evolution process. Further analyses of gene duplication events showed that whole-genome duplication, transposed duplication, and dispersed duplication events were the primary forces driving the R2R3-MYB gene family expansion of Ipomoea plants, and these duplicated genes experienced strong purifying selection because of their Ka/Ks ratio, which is less than 1. Additionally, the genomic sequence length of 131 IbR2R3-MYBs varied from 923 bp to ~12.9 kb with a mean of ~2.6 kb, and most of them had more than three exons. The Motif 1, 2, 3, and 4 formed typical R2 and R3 domains and were identified in all IbR2R3-MYB proteins. Finally, based on multiple RNA-seq datasets, two IbR2R3-MYB genes (IbMYB1/g17138.t1 and IbMYB113/g17108.t1) were relatively highly expressed in pigmented leaves and tuberous root flesh and skin, respectively; thus, they were identified to regulate tissue-specific anthocyanin accumulation in sweet potato. This study provides a basis for the evolution and function of the R2R3-MYB gene family in sweet potatoes and five other Ipomoea species.
Keywords: Ipomoea species; R2R3-MYB gene family; anthocyanin biosynthesis; expression analysis; sweet potato.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures





Similar articles
-
Comparative analysis of the MYB gene family in seven Ipomoea species.Front Plant Sci. 2023 Mar 20;14:1155018. doi: 10.3389/fpls.2023.1155018. eCollection 2023. Front Plant Sci. 2023. PMID: 37021302 Free PMC article.
-
The Expression of IbMYB1 Is Essential to Maintain the Purple Color of Leaf and Storage Root in Sweet Potato [Ipomoea batatas (L.) Lam].Front Plant Sci. 2021 Sep 23;12:688707. doi: 10.3389/fpls.2021.688707. eCollection 2021. Front Plant Sci. 2021. PMID: 34630449 Free PMC article.
-
Overexpression of the IbMYB1 gene in an orange-fleshed sweet potato cultivar produces a dual-pigmented transgenic sweet potato with improved antioxidant activity.Physiol Plant. 2015 Apr;153(4):525-37. doi: 10.1111/ppl.12281. Epub 2014 Oct 28. Physiol Plant. 2015. PMID: 25220246
-
Genome-wide identification of Pistacia R2R3-MYB gene family and function characterization of PcMYB113 during autumn leaf coloration in Pistacia chinensis.Int J Biol Macromol. 2021 Dec 1;192:16-27. doi: 10.1016/j.ijbiomac.2021.09.092. Epub 2021 Sep 20. Int J Biol Macromol. 2021. PMID: 34555399 Review.
-
Current status in whole genome sequencing and analysis of Ipomoea spp.Plant Cell Rep. 2019 Nov;38(11):1365-1371. doi: 10.1007/s00299-019-02464-4. Epub 2019 Aug 29. Plant Cell Rep. 2019. PMID: 31468128 Free PMC article. Review.
Cited by
-
In silico analysis of R2R3-MYB transcription factors in the basal eudicot model, Aquilegia coerulea.3 Biotech. 2024 Nov;14(11):284. doi: 10.1007/s13205-024-04119-y. Epub 2024 Oct 29. 3 Biotech. 2024. PMID: 39479299 Free PMC article.
-
Integration of comparative transcriptomics and WGCNA characterizes the regulation of anthocyanin biosynthesis in mung bean (Vigna radiata L.).Front Plant Sci. 2023 Oct 24;14:1251464. doi: 10.3389/fpls.2023.1251464. eCollection 2023. Front Plant Sci. 2023. PMID: 37941672 Free PMC article.
-
Transcription factors in Orinus: novel insights into transcription regulation for speciation adaptation on the Qinghai-Xizang (Tibet) Plateau.BMC Plant Biol. 2025 Apr 29;25(1):560. doi: 10.1186/s12870-025-06602-x. BMC Plant Biol. 2025. PMID: 40301765 Free PMC article.
References
-
- FAOSTAT Food and Agricultural Organization from the United Nations. [(accessed on 14 April 2023)]. Available online: http://www.fao.org/faostat/en/#data/QC.
-
- Mohanraj R. Sweet Potato: Bioactive Compounds and Health Benefits. In: Mérillon J.-M., Ramawat K.G., editors. Bioactive Molecules in Food. Springer International Publishing; Cham, Switerland: 2018. pp. 1–16. (Reference Series in Phytochemistry).
Grants and funding
LinkOut - more resources
Full Text Sources

