Development of a Pan- Filoviridae SYBR Green qPCR Assay for Biosurveillance Studies in Bats
- PMID: 37112966
- PMCID: PMC10145118
- DOI: 10.3390/v15040987
Development of a Pan- Filoviridae SYBR Green qPCR Assay for Biosurveillance Studies in Bats
Abstract
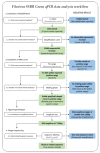
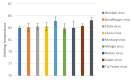
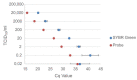
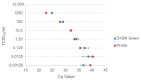
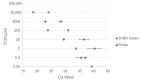
Recent studies have indicated that bats are hosts to diverse filoviruses. Currently, no pan-filovirus molecular assays are available that have been evaluated for the detection of all mammalian filoviruses. In this study, a two-step pan-filovirus SYBR Green real-time PCR assay targeting the nucleoprotein gene was developed for filovirus surveillance in bats. Synthetic constructs were designed as representatives of nine filovirus species and used to evaluate the assay. This assay detected all synthetic constructs included with an analytical sensitivity of 3-31.7 copies/reaction and was evaluated against the field collected samples. The assay's performance was similar to a previously published probe based assay for detecting Ebola- and Marburgvirus. The developed pan-filovirus SYBR Green assay will allow for more affordable and sensitive detection of mammalian filoviruses in bat samples.
Keywords: SYBR Green; bats; pan-filovirus; qPCR; surveillance.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Development, validation and clinical evaluation of a broad-range pan-filovirus RT-qPCR.J Clin Virol. 2019 May;114:26-31. doi: 10.1016/j.jcv.2019.03.010. Epub 2019 Mar 19. J Clin Virol. 2019. PMID: 30904708
-
Detection of filovirus-reactive antibodies in Australian bat species.J Gen Virol. 2022 Aug;103(8). doi: 10.1099/jgv.0.001785. J Gen Virol. 2022. PMID: 35972225
-
Broad and Temperature Independent Replication Potential of Filoviruses on Cells Derived From Old and New World Bat Species.J Infect Dis. 2016 Oct 15;214(suppl 3):S297-S302. doi: 10.1093/infdis/jiw199. Epub 2016 Jun 26. J Infect Dis. 2016. PMID: 27354372 Free PMC article.
-
Filoviruses in bats: current knowledge and future directions.Viruses. 2014 Apr 17;6(4):1759-88. doi: 10.3390/v6041759. Viruses. 2014. PMID: 24747773 Free PMC article. Review.
-
ICTV Virus Taxonomy Profile: Filoviridae.J Gen Virol. 2019 Jun;100(6):911-912. doi: 10.1099/jgv.0.001252. Epub 2019 Apr 25. J Gen Virol. 2019. PMID: 31021739 Free PMC article. Review.
Cited by
-
Identification and Characterization of Viral and Bacterial Pathogens in Free-Living Bats of Kopaonik National Park, Serbia.Vet Sci. 2025 Apr 24;12(5):401. doi: 10.3390/vetsci12050401. Vet Sci. 2025. PMID: 40431494 Free PMC article.
-
A high-throughput, polymerase-targeted RT-PCR for broad detection of mammalian filoviruses.Microbiol Spectr. 2024 Sep 3;12(9):e0101024. doi: 10.1128/spectrum.01010-24. Epub 2024 Jul 24. Microbiol Spectr. 2024. PMID: 39046245 Free PMC article.
References
-
- Kuhn J.H., Adkins S., Agwanda B.R., Al Kubrusli R., Alkhovsky S.V., Amarasinghe G.K., Avšič-Županc T., Ayllón M.A., Bahl J., Balkema-Buschmann A., et al. 2021 Taxonomic Update of Phylum Negarnaviricota (Riboviria: Orthornavirae), Including the Large Orders Bunyavirales and Mononegavirales. Arch. Virol. 2021;166:3513–3566. doi: 10.1007/s00705-021-05143-6. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

