RAPID prep: A Simple, Fast Protocol for RNA Metagenomic Sequencing of Clinical Samples
- PMID: 37112986
- PMCID: PMC10146689
- DOI: 10.3390/v15041006
RAPID prep: A Simple, Fast Protocol for RNA Metagenomic Sequencing of Clinical Samples
Abstract
Emerging infectious disease threats require rapid response tools to inform diagnostics, treatment, and outbreak control. RNA-based metagenomics offers this; however, most approaches are time-consuming and laborious. Here, we present a simple and fast protocol, the RAPIDprep assay, with the aim of providing a cause-agnostic laboratory diagnosis of infection within 24 h of sample collection by sequencing ribosomal RNA-depleted total RNA. The method is based on the synthesis and amplification of double-stranded cDNA followed by short-read sequencing, with minimal handling and clean-up steps to improve processing time. The approach was optimized and applied to a range of clinical respiratory samples to demonstrate diagnostic and quantitative performance. Our results showed robust depletion of both human and microbial rRNA, and library amplification across different sample types, qualities, and extraction kits using a single workflow without input nucleic-acid quantification or quality assessment. Furthermore, we demonstrated the genomic yield of both known and undiagnosed pathogens with complete genomes recovered in most cases to inform molecular epidemiological investigations and vaccine design. The RAPIDprep assay is a simple and effective tool, and representative of an important shift toward the integration of modern genomic techniques with infectious disease investigations.
Keywords: RNA sequencing; diagnostics; infectious diseases; metagenomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

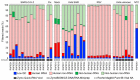

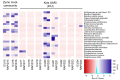
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

