Complement C1q-dependent excitatory and inhibitory synapse elimination by astrocytes and microglia in Alzheimer's disease mouse models
- PMID: 37118504
- PMCID: PMC10154216
- DOI: 10.1038/s43587-022-00281-1
Complement C1q-dependent excitatory and inhibitory synapse elimination by astrocytes and microglia in Alzheimer's disease mouse models
Abstract
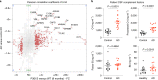
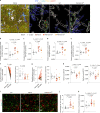
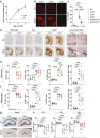
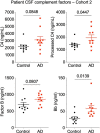
Microglia and complement can mediate neurodegeneration in Alzheimer's disease (AD). By integrative multi-omics analysis, here we show that astrocytic and microglial proteins are increased in TauP301S synapse fractions with age and in a C1q-dependent manner. In addition to microglia, we identified that astrocytes contribute substantially to synapse elimination in TauP301S hippocampi. Notably, we found relatively more excitatory synapse marker proteins in astrocytic lysosomes, whereas microglial lysosomes contained more inhibitory synapse material. C1q deletion reduced astrocyte-synapse association and decreased astrocytic and microglial synapses engulfment in TauP301S mice and rescued synapse density. Finally, in an AD mouse model that combines β-amyloid and Tau pathologies, deletion of the AD risk gene Trem2 impaired microglial phagocytosis of synapses, whereas astrocytes engulfed more inhibitory synapses around plaques. Together, our data reveal that astrocytes contact and eliminate synapses in a C1q-dependent manner and thereby contribute to pathological synapse loss and that astrocytic phagocytosis can compensate for microglial dysfunction.
© 2022. The Author(s).
Conflict of interest statement
Several authors were full-time employees of Genentech during the course of the study. M.S. is scientific co-founder and member of the SAB of Neumora Therapeutics and member of the SAB of Vanqua Bio, ArcLight Therapeutics and Cerevel Therapeutics. All other authors declare no competing interests.
Figures









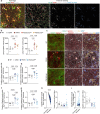
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

