Improving Yields in Multi-analyte Extractions by Utilizing Post-homogenized Tissue Debris
- PMID: 37119238
- PMCID: PMC10227881
- DOI: 10.1369/00221554231172823
Improving Yields in Multi-analyte Extractions by Utilizing Post-homogenized Tissue Debris
Abstract
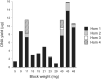
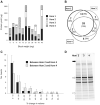
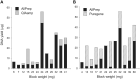
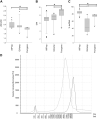
In multi-analyte extractions, tissue is typically homogenized in a lysis buffer, and then DNA, RNA, and protein are purified from the supernatant. However, yields are typically lower than in dedicated, single-analyte extractions. In a two-part experiment, we assessed whether yields could be improved by revisiting the normally discarded, post-homogenized tissue debris. We initially performed additional homogenizations, each followed by a simultaneous extraction. These yielded no additional RNA, 13% additional DNA (which became progressively more degraded), and 161.7% additional protein (which changed in proteome when analyzed using SDS-PAGE). We then digested post-homogenized tissue debris from a simultaneous extraction using proteinase K and extracted DNA using silica spin columns or alcohol precipitation. An average additional DNA yield of 27.1% (silica spin columns) or 203.9% (alcohol precipitation) was obtained with/without compromising DNA integrity (assessment by long-range PCR, DNA Integrity Numbers, and size at peak fluorescence of electropherogram). Validation using a cohort of 65 tissue blocks returned an average additional DNA yield of 31.6% (silica columns) and 54.8% (alcohol precipitation). Users can therefore refreeze the homogenized remnants of tissue blocks rather than disposing of them and then perform additional DNA extractions if yields in the initial multi-analyte extractions were low.
Keywords: AllPrep; Puregene; QIAamp; biospecimen science; homogenization; optimization; simultaneous.
Conflict of interest statement
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Figures
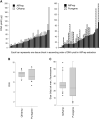
References
-
- Mathieson W, Thomas GA. Simultaneously extracting DNA, RNA, and protein using kits: is sample quantity or quality prejudiced? Anal Biochem. 2013;433(1):10–8. - PubMed
-
- Skiadas J, Aston C, Samad A, Anantharaman TS, Mishra B, Schwartz DC. Optical PCR: genomic analysis by long-range PCR and optical mapping. Mamm Genome. 1999;10(10):1005–9. - PubMed
-
- Mathieson W, Sanchez I, Mommaerts K, Frasquilho S, Betsou F. An independent evaluation of the CryoXtract Instruments’ CXT350 frozen sample aliquotter using tissue and fecal biospecimens. Biopreserv Biobank. 2016;14:2–8. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

