SARS-CoV-2 infection and recovery in children: Distinct T cell responses in MIS-C compared to COVID-19
- PMID: 37133746
- PMCID: PMC10163842
- DOI: 10.1084/jem.20221518
SARS-CoV-2 infection and recovery in children: Distinct T cell responses in MIS-C compared to COVID-19
Abstract
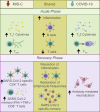
SARS-CoV-2 infection for most children results in mild or minimal symptoms, though in rare cases severe disease can develop, including a multisystem inflammatory syndrome (MIS-C) with myocarditis. Here, we present longitudinal profiling of immune responses during acute disease and following recovery in children who developed MIS-C, relative to children who experienced more typical symptoms of COVID-19. T cells in acute MIS-C exhibited transient signatures of activation, inflammation, and tissue residency which correlated with cardiac disease severity, while T cells in acute COVID-19 upregulated markers of follicular helper T cells for promoting antibody production. The resultant memory immune response in recovery showed increased frequencies of virus-specific memory T cells with pro-inflammatory functions in children with prior MIS-C compared to COVID-19 while both cohorts generated comparable antibody responses. Together our results reveal distinct effector and memory T cell responses in pediatric SARS-CoV-2 infection delineated by clinical syndrome, and a potential role for tissue-derived T cells in the immune pathology of systemic disease.
© 2023 Rybkina et al.
Conflict of interest statement
Disclosures: B.R. Anderson reported grants from Genentech and the National Institutes of Health outside the submitted work. J. Milner reported personal fees from Blueprint Medicines outside the submitted work. T.J. Connors reported grants from the National Institutes of Health during the conduct of the study and outside the submitted work. No other disclosures were reported.
Figures








Similar articles
-
Long-term humoral signatures following acute pediatric COVID-19 and Multisystem Inflammatory Syndrome in Children.Pediatr Res. 2023 Oct;94(4):1327-1334. doi: 10.1038/s41390-023-02627-w. Epub 2023 May 12. Pediatr Res. 2023. PMID: 37173406 Free PMC article.
-
SARS-CoV-2-specific T cell responses in patients with multisystem inflammatory syndrome in children.Clin Immunol. 2022 Oct;243:109106. doi: 10.1016/j.clim.2022.109106. Epub 2022 Aug 30. Clin Immunol. 2022. PMID: 36049601 Free PMC article.
-
Limited induction of SARS-CoV-2-specific T cell responses in children with multisystem inflammatory syndrome compared with COVID-19.JCI Insight. 2022 Feb 22;7(4):e155145. doi: 10.1172/jci.insight.155145. JCI Insight. 2022. PMID: 35044955 Free PMC article.
-
Current Understanding of Multisystem Inflammatory Syndrome (MIS-C) Following COVID-19 and Its Distinction from Kawasaki Disease.Curr Rheumatol Rep. 2021 Jul 3;23(8):58. doi: 10.1007/s11926-021-01028-4. Curr Rheumatol Rep. 2021. PMID: 34216296 Free PMC article. Review.
-
Multisystem Inflammatory Syndrome in Children (MIS-C).Curr Allergy Asthma Rep. 2022 May;22(5):53-60. doi: 10.1007/s11882-022-01031-4. Epub 2022 Mar 22. Curr Allergy Asthma Rep. 2022. PMID: 35314921 Free PMC article. Review.
Cited by
-
Durable T cell immunity to COVID-19 vaccines in MS patients on B cell depletion therapy.NPJ Vaccines. 2025 May 17;10(1):98. doi: 10.1038/s41541-025-01151-8. NPJ Vaccines. 2025. PMID: 40382362 Free PMC article.
-
Liver abnormalities following SARS-CoV-2 infection in children 1 to 10 years of age.Fam Med Community Health. 2024 Jan 25;12(1):e002655. doi: 10.1136/fmch-2023-002655. Fam Med Community Health. 2024. PMID: 38272541 Free PMC article.
-
Molecular mimicry in multisystem inflammatory syndrome in children.Nature. 2024 Aug;632(8025):622-629. doi: 10.1038/s41586-024-07722-4. Epub 2024 Aug 7. Nature. 2024. PMID: 39112696 Free PMC article.
-
Peripheral immune progression to long COVID is associated with mitochondrial gene transcription: A meta-analysis.Mitochondrion. 2025 Jul 28;85:102072. doi: 10.1016/j.mito.2025.102072. Online ahead of print. Mitochondrion. 2025. PMID: 40738238 Free PMC article.
-
Distinct Localization, Transcriptional Profiles, and Functionality in Early Life Tonsil Regulatory T Cells.J Immunol. 2024 Aug 1;213(3):306-316. doi: 10.4049/jimmunol.2300890. J Immunol. 2024. PMID: 38905110 Free PMC article.
References
-
- Belhadjer, Z., Auriau J., Méot M., Oualha M., Renolleau S., Houyel L., and Bonnet D.. 2020. Addition of corticosteroids to immunoglobulins is associated with recovery of cardiac function in multi-inflammatory syndrome in children. Circulation. 142:2282–2284. 10.1161/CIRCULATIONAHA.120.050147 - DOI - PMC - PubMed
Publication types
MeSH terms
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

