Assembly properties of bacterial actin MreB involved in Spiroplasma swimming motility
- PMID: 37150324
- PMCID: PMC10279915
- DOI: 10.1016/j.jbc.2023.104793
Assembly properties of bacterial actin MreB involved in Spiroplasma swimming motility
Abstract
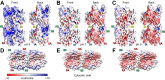
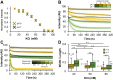
Bacterial actin MreB forms filaments composed of antiparallel double-stranded units. The wall-less helical bacterium Spiroplasma has five MreB homologs (MreB1-5), some of which are involved in an intracellular ribbon for driving the bacterium's swimming motility. Although the interaction between MreB units is important for understanding Spiroplasma swimming, the interaction modes of each ribbon component are unclear. Here, we examined the assembly properties of Spiroplasma eriocheiris MreB5 (SpeMreB5), one of the ribbon component proteins that forms sheets. Electron microscopy revealed that sheet formation was inhibited under acidic conditions and bundle structures were formed under acidic and neutral conditions with low ionic strength. We also used solution assays and identified four properties of SpeMreB5 bundles as follows: (I) bundle formation followed sheet formation; (II) electrostatic interactions were required for bundle formation; (III) the positively charged and unstructured C-terminal region contributed to promoting lateral interactions for bundle formation; and (IV) bundle formation required Mg2+ at neutral pH but was inhibited by divalent cations under acidic pH conditions. During these studies, we also characterized two aggregation modes of SpeMreB5 with distinct responses to ATP. These properties will shed light on SpeMreB5 assembly dynamics at the molecular level.
Keywords: ATPase; Mollicutes; Mycoplasma; actin; bacteria; bundle formation; cell motility; cytoskeleton; electron microscopy; polymerization dynamics.
Copyright © 2023 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures





Similar articles
-
A bacterial actin with high ATPase activity regulates the polymerization of a partner MreB isoform essential for Spiroplasma swimming motility.J Biol Chem. 2025 Jul 7;301(8):110462. doi: 10.1016/j.jbc.2025.110462. Online ahead of print. J Biol Chem. 2025. PMID: 40633615 Free PMC article.
-
ATP-dependent polymerization dynamics of bacterial actin proteins involved in Spiroplasma swimming.Open Biol. 2022 Oct;12(10):220083. doi: 10.1098/rsob.220083. Epub 2022 Oct 26. Open Biol. 2022. PMID: 36285441 Free PMC article.
-
Purification and ATPase Activity Measurement of Spiroplasma MreB.Methods Mol Biol. 2023;2646:359-371. doi: 10.1007/978-1-0716-3060-0_30. Methods Mol Biol. 2023. PMID: 36842130
-
Shaping and moving a spiroplasma.J Mol Microbiol Biotechnol. 2004;7(1-2):78-87. doi: 10.1159/000077872. J Mol Microbiol Biotechnol. 2004. PMID: 15170406 Review.
-
Molecules of the bacterial cytoskeleton.Annu Rev Biophys Biomol Struct. 2004;33:177-98. doi: 10.1146/annurev.biophys.33.110502.132647. Annu Rev Biophys Biomol Struct. 2004. PMID: 15139810 Review.
Cited by
-
Deciphering the impact of MreB on the morphology and pathogenicity of the aquatic pathogen Spiroplasma eriocheiris.Biol Direct. 2024 Oct 23;19(1):98. doi: 10.1186/s13062-024-00537-3. Biol Direct. 2024. PMID: 39444023 Free PMC article.
-
MreB: unraveling the molecular mechanisms of bacterial shape, division, and environmental adaptation.Cell Commun Signal. 2025 Aug 22;23(1):377. doi: 10.1186/s12964-025-02373-y. Cell Commun Signal. 2025. PMID: 40847400 Free PMC article. Review.
-
A bacterial actin with high ATPase activity regulates the polymerization of a partner MreB isoform essential for Spiroplasma swimming motility.J Biol Chem. 2025 Jul 7;301(8):110462. doi: 10.1016/j.jbc.2025.110462. Online ahead of print. J Biol Chem. 2025. PMID: 40633615 Free PMC article.
References
-
- Wagstaff J., Löwe J. Prokaryotic cytoskeletons: protein filaments organizing small cells. Nat. Rev. Microbiol. 2018;16:187–201. - PubMed
-
- Harne S., Duret S., Pande V., Bapat M., Béven L., Gayathri P. MreB5 is a determinant of rod-to-helical transition in the cell-wall-less bacterium Spiroplasma. Curr. Biol. 2020;30:4753–4762.e7. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources

