A deep learning algorithm to predict risk of pancreatic cancer from disease trajectories
- PMID: 37156936
- PMCID: PMC10202814
- DOI: 10.1038/s41591-023-02332-5
A deep learning algorithm to predict risk of pancreatic cancer from disease trajectories
Abstract
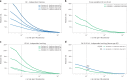
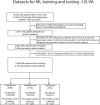
Pancreatic cancer is an aggressive disease that typically presents late with poor outcomes, indicating a pronounced need for early detection. In this study, we applied artificial intelligence methods to clinical data from 6 million patients (24,000 pancreatic cancer cases) in Denmark (Danish National Patient Registry (DNPR)) and from 3 million patients (3,900 cases) in the United States (US Veterans Affairs (US-VA)). We trained machine learning models on the sequence of disease codes in clinical histories and tested prediction of cancer occurrence within incremental time windows (CancerRiskNet). For cancer occurrence within 36 months, the performance of the best DNPR model has area under the receiver operating characteristic (AUROC) curve = 0.88 and decreases to AUROC (3m) = 0.83 when disease events within 3 months before cancer diagnosis are excluded from training, with an estimated relative risk of 59 for 1,000 highest-risk patients older than age 50 years. Cross-application of the Danish model to US-VA data had lower performance (AUROC = 0.71), and retraining was needed to improve performance (AUROC = 0.78, AUROC (3m) = 0.76). These results improve the ability to design realistic surveillance programs for patients at elevated risk, potentially benefiting lifespan and quality of life by early detection of this aggressive cancer.
© 2023. The Author(s).
Conflict of interest statement
S.B. has ownership in Intomics A/S, Hoba Therapeutics Aps, Novo Nordisk A/S, Lundbeck A/S and ALK Abello and has managing board memberships in Proscion A/S and Intomics A/S. B.M.W. notes grant funding from Celgene and Eli Lilly and consulting fees from BioLineRx, Celgene and GRAIL. A.R. is a co-founder and equity holder of Celsius Therapeutics, an equity holder in Immunitas and was a scientific advisory board member of Thermo Fisher Scientific, Syros Pharmaceuticals, Neogene Therapeutics and Asimov until 31 July 2020. D.S.M. is an advisor for Dyno Therapeutics, Octant, Jura Bio, Tectonic Therapeutic and Genentech and is a co-founder of Seismic Therapeutic. C.S. is on the scientific advisory board of CytoReason. From 1 August 2020, A.R. is an employee of Genentech and has equity in Roche. The remaining authors declare no competing interests.
Figures





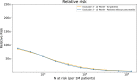

References
-
- Rahib L, et al. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014;74:2913–2921. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

