Pathobionts in the tumour microbiota predict survival following resection for colorectal cancer
- PMID: 37158960
- PMCID: PMC10165813
- DOI: 10.1186/s40168-023-01518-w
Pathobionts in the tumour microbiota predict survival following resection for colorectal cancer
Abstract
Background and aims: The gut microbiota is implicated in the pathogenesis of colorectal cancer (CRC). We aimed to map the CRC mucosal microbiota and metabolome and define the influence of the tumoral microbiota on oncological outcomes.
Methods: A multicentre, prospective observational study was conducted of CRC patients undergoing primary surgical resection in the UK (n = 74) and Czech Republic (n = 61). Analysis was performed using metataxonomics, ultra-performance liquid chromatography-mass spectrometry (UPLC-MS), targeted bacterial qPCR and tumour exome sequencing. Hierarchical clustering accounting for clinical and oncological covariates was performed to identify clusters of bacteria and metabolites linked to CRC. Cox proportional hazards regression was used to ascertain clusters associated with disease-free survival over median follow-up of 50 months.
Results: Thirteen mucosal microbiota clusters were identified, of which five were significantly different between tumour and paired normal mucosa. Cluster 7, containing the pathobionts Fusobacterium nucleatum and Granulicatella adiacens, was strongly associated with CRC (PFDR = 0.0002). Additionally, tumoral dominance of cluster 7 independently predicted favourable disease-free survival (adjusted p = 0.031). Cluster 1, containing Faecalibacterium prausnitzii and Ruminococcus gnavus, was negatively associated with cancer (PFDR = 0.0009), and abundance was independently predictive of worse disease-free survival (adjusted p = 0.0009). UPLC-MS analysis revealed two major metabolic (Met) clusters. Met 1, composed of medium chain (MCFA), long-chain (LCFA) and very long-chain (VLCFA) fatty acid species, ceramides and lysophospholipids, was negatively associated with CRC (PFDR = 2.61 × 10-11); Met 2, composed of phosphatidylcholine species, nucleosides and amino acids, was strongly associated with CRC (PFDR = 1.30 × 10-12), but metabolite clusters were not associated with disease-free survival (p = 0.358). An association was identified between Met 1 and DNA mismatch-repair deficiency (p = 0.005). FBXW7 mutations were only found in cancers predominant in microbiota cluster 7.
Conclusions: Networks of pathobionts in the tumour mucosal niche are associated with tumour mutation and metabolic subtypes and predict favourable outcome following CRC resection. Video Abstract.
Keywords: Colorectal cancer; Gut microbiota; Metabolome; Metataxonomics.
© 2023. The Author(s).
Conflict of interest statement
AD is Chair of the Health Security initiative at Flagship Pioneering UK Ltd. DC declares being on the scientific advisory board for OVIBIO, and grant funding from MedImmune, Clovis, Eli Lilly, 4SC, Bayer, Celgene, Leap & Roche, all made payable to the Royal Marsden Hospital.
Figures

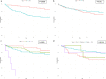


References
-
- Cancer Research UK: Bowel cancer statistics https://www.cancerresearchuk.org/health-professional/cancer-statistics/s... Accessed August 2020.
-
- Partnership HQI. National Bowel Cancer Audit Annual Report 2020.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

