The T-cell-directed vaccine BNT162b4 encoding conserved non-spike antigens protects animals from severe SARS-CoV-2 infection
- PMID: 37164012
- PMCID: PMC10099181
- DOI: 10.1016/j.cell.2023.04.007
The T-cell-directed vaccine BNT162b4 encoding conserved non-spike antigens protects animals from severe SARS-CoV-2 infection
Abstract
T cell responses play an important role in protection against beta-coronavirus infections, including SARS-CoV-2, where they associate with decreased COVID-19 disease severity and duration. To enhance T cell immunity across epitopes infrequently altered in SARS-CoV-2 variants, we designed BNT162b4, an mRNA vaccine component that is intended to be combined with BNT162b2, the spike-protein-encoding vaccine. BNT162b4 encodes variant-conserved, immunogenic segments of the SARS-CoV-2 nucleocapsid, membrane, and ORF1ab proteins, targeting diverse HLA alleles. BNT162b4 elicits polyfunctional CD4+ and CD8+ T cell responses to diverse epitopes in animal models, alone or when co-administered with BNT162b2 while preserving spike-specific immunity. Importantly, we demonstrate that BNT162b4 protects hamsters from severe disease and reduces viral titers following challenge with viral variants. These data suggest that a combination of BNT162b2 and BNT162b4 could reduce COVID-19 disease severity and duration caused by circulating or future variants. BNT162b4 is currently being clinically evaluated in combination with the BA.4/BA.5 Omicron-updated bivalent BNT162b2 (NCT05541861).
Keywords: COVID-19; HLA ligandomics; SARS-CoV-2; T cell responses; immune protection; mRNA T cell vaccine; vaccine design; viral variants.
Copyright © 2023 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests U.S. and Ö.T. are management board members and employees at BioNTech SE. R.B.G., Board of Directors, Alkermes plc, Infinity Pharmaceuticals, and Zai Laboratory, and Scientific Advisory Board, Leap Therapeutics; consultant Third Rock Ventures, stockholder and employee of BioNTech US. C.M.A., Y.J.X., D.A.R., H.D., D.H., S.M., K.M., B.K., T.E.S., A.L., A.Z., S.A.K., C.I.C.C., S.H., A.P.H., T.Z., Y.L.-L., K.W., A.B.V., J.R.S., S.F., K.T., K.W., T.A.A., Ö.T., U.S., R.B.G., and A.P. are either current or past employees of BioNTech SE or BioNTech US, are stockholders, and/or are inventors on patents and patent applications related to RNA technology and COVID-19 vaccines.
Figures


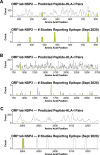











References
-
- Muik, A., Lui, B.G., Diao, H., Fu, Y., Bacher, M., Toker, A., Grosser, J., Ozhelvaci, O., Grikscheit, K., Hoehl, S., et al. (2022). Progressive Loss of Conserved Spike Protein Neutralizing Antibody Sites in Omicron Sublineages Is Balanced by Preserved T-Cell Recognition Epitopes. bioRxiv, 2022.2012.2015.520569. 10.1101/2022.12.15.520569
MeSH terms
Substances
Supplementary concepts
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

