Identification of a genomic DNA sequence that quantitatively modulates KLF1 transcription factor expression in differentiating human hematopoietic cells
- PMID: 37165057
- PMCID: PMC10172341
- DOI: 10.1038/s41598-023-34805-5
Identification of a genomic DNA sequence that quantitatively modulates KLF1 transcription factor expression in differentiating human hematopoietic cells
Abstract
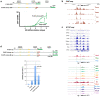
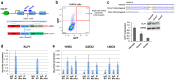
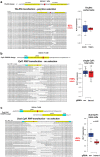
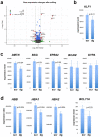
The onset of erythropoiesis is under strict developmental control, with direct and indirect inputs influencing its derivation from the hematopoietic stem cell. A major regulator of this transition is KLF1/EKLF, a zinc finger transcription factor that plays a global role in all aspects of erythropoiesis. Here, we have identified a short, conserved enhancer element in KLF1 intron 1 that is important for establishing optimal levels of KLF1 in mouse and human cells. Chromatin accessibility of this site exhibits cell-type specificity and is under developmental control during the differentiation of human CD34+ cells towards the erythroid lineage. This site binds GATA1, SMAD1, TAL1, and ETV6. In vivo editing of this region in cell lines and primary cells reduces KLF1 expression quantitatively. However, we find that, similar to observations seen in pedigrees of families with KLF1 mutations, downstream effects are variable, suggesting that the global architecture of the site is buffered towards keeping the KLF1 genetic region in an active state. We propose that modification of intron 1 in both alleles is not equivalent to complete loss of function of one allele.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Erythroid Krüppel-Like Factor (KLF1): A Surprisingly Versatile Regulator of Erythroid Differentiation.Adv Exp Med Biol. 2024;1459:217-242. doi: 10.1007/978-3-031-62731-6_10. Adv Exp Med Biol. 2024. PMID: 39017846 Free PMC article. Review.
-
Ldb1 complexes: the new master regulators of erythroid gene transcription.Trends Genet. 2014 Jan;30(1):1-9. doi: 10.1016/j.tig.2013.10.001. Epub 2013 Nov 27. Trends Genet. 2014. PMID: 24290192 Free PMC article. Review.
-
A Positive Regulatory Feedback Loop between EKLF/KLF1 and TAL1/SCL Sustaining the Erythropoiesis.Int J Mol Sci. 2021 Jul 27;22(15):8024. doi: 10.3390/ijms22158024. Int J Mol Sci. 2021. PMID: 34360789 Free PMC article.
-
Novel roles for KLF1 in erythropoiesis revealed by mRNA-seq.Genome Res. 2012 Dec;22(12):2385-98. doi: 10.1101/gr.135707.111. Epub 2012 Jul 26. Genome Res. 2012. PMID: 22835905 Free PMC article.
-
EKLF/KLF1 controls cell cycle entry via direct regulation of E2f2.J Biol Chem. 2009 Jul 31;284(31):20966-74. doi: 10.1074/jbc.M109.006346. Epub 2009 May 20. J Biol Chem. 2009. PMID: 19457859 Free PMC article.
Cited by
-
Key Genes FECH and ALAS2 under Acute High-Altitude Exposure: A Gene Expression and Network Analysis Based on Expression Profile Data.Genes (Basel). 2024 Aug 14;15(8):1075. doi: 10.3390/genes15081075. Genes (Basel). 2024. PMID: 39202434 Free PMC article.
-
Association of DDX5/p68 protein with the upstream erythroid enhancer element (EHS1) of the gene encoding the KLF1 transcription factor.J Biol Chem. 2023 Dec;299(12):105489. doi: 10.1016/j.jbc.2023.105489. Epub 2023 Nov 22. J Biol Chem. 2023. PMID: 38000658 Free PMC article.
-
Erythroid Krüppel-Like Factor (KLF1): A Surprisingly Versatile Regulator of Erythroid Differentiation.Adv Exp Med Biol. 2024;1459:217-242. doi: 10.1007/978-3-031-62731-6_10. Adv Exp Med Biol. 2024. PMID: 39017846 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

