Precise targeting for 3D cryo-correlative light and electron microscopy volume imaging of tissues using a FinderTOP
- PMID: 37169904
- PMCID: PMC10175257
- DOI: 10.1038/s42003-023-04887-y
Precise targeting for 3D cryo-correlative light and electron microscopy volume imaging of tissues using a FinderTOP
Abstract
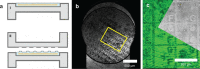
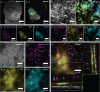
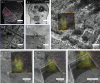
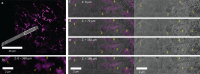
Cryo-correlative light and electron microscopy (cryoCLEM) is a powerful strategy to high resolution imaging in the unperturbed hydrated state. In this approach fluorescence microscopy aids localizing the area of interest, and cryogenic focused ion beam/scanning electron microscopy (cryoFIB/SEM) allows preparation of thin cryo-lamellae for cryoET. However, the current method cannot be accurately applied on bulky (3D) samples such as tissues and organoids. 3D cryo-correlative imaging of large volumes is needed to close the resolution gap between cryo-light microscopy and cryoET, placing sub-nanometer observations in a larger biological context. Currently technological hurdles render 3D cryoCLEM an unexplored approach. Here we demonstrate a cryoCLEM workflow for tissues, correlating cryo-Airyscan confocal microscopy with 3D cryoFIB/SEM volume imaging. Accurate correlation is achieved by imprinting a FinderTOP pattern in the sample surface during high pressure freezing, and allows precise targeting for cryoFIB/SEM volume imaging.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Correlative cryo-microscopy pipelines for in situ cellular studies.Methods Cell Biol. 2024;187:175-203. doi: 10.1016/bs.mcb.2024.02.038. Epub 2024 Mar 27. Methods Cell Biol. 2024. PMID: 38705624
-
Multi-scale 3D Cryo-Correlative Microscopy for Vitrified Cells.Structure. 2020 Nov 3;28(11):1231-1237.e3. doi: 10.1016/j.str.2020.07.017. Epub 2020 Aug 18. Structure. 2020. PMID: 32814034 Free PMC article.
-
High-precision correlative fluorescence and electron cryo microscopy using two independent alignment markers.Ultramicroscopy. 2014 Aug;143(100):41-51. doi: 10.1016/j.ultramic.2013.10.011. Epub 2013 Oct 19. Ultramicroscopy. 2014. PMID: 24262358 Free PMC article.
-
Correlative cryo-electron tomography and optical microscopy of cells.Curr Opin Struct Biol. 2013 Oct;23(5):763-70. doi: 10.1016/j.sbi.2013.07.017. Epub 2013 Aug 17. Curr Opin Struct Biol. 2013. PMID: 23962486 Free PMC article. Review.
-
Advances in Cryo-Correlative Light and Electron Microscopy: Applications for Studying Molecular and Cellular Events.Protein J. 2019 Dec;38(6):609-615. doi: 10.1007/s10930-019-09856-1. Protein J. 2019. PMID: 31396855 Review.
Cited by
-
Unraveling atomic complexity from frozen samples.Struct Dyn. 2025 Apr 18;12(2):020901. doi: 10.1063/4.0000303. eCollection 2025 Mar. Struct Dyn. 2025. PMID: 40255534 Free PMC article.
-
Horizontal gene transfer and beyond: the delivery of biological matter by bacterial membrane vesicles to host and bacterial cells.Curr Opin Microbiol. 2024 Oct;81:102525. doi: 10.1016/j.mib.2024.102525. Epub 2024 Aug 26. Curr Opin Microbiol. 2024. PMID: 39190937 Review.
-
Cryo-electron microscopy in the study of virus entry and infection.Front Mol Biosci. 2024 Jul 24;11:1429180. doi: 10.3389/fmolb.2024.1429180. eCollection 2024. Front Mol Biosci. 2024. PMID: 39114367 Free PMC article. Review.
References
-
- The Nobel Prize in Chemistry 2017. NobelPrize.org. Nobel Prize Outreach AB 2022. 2022. https://www.nobelprize.org/prizes/chemistry/2017/summary/.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

