Influenza A virus exploits transferrin receptor recycling to enter host cells
- PMID: 37192162
- PMCID: PMC10214170
- DOI: 10.1073/pnas.2214936120
Influenza A virus exploits transferrin receptor recycling to enter host cells
Abstract
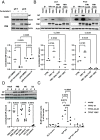
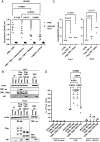
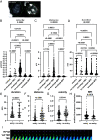
Influenza A virus (IAV) enters host cells mostly through clathrin-dependent receptor-mediated endocytosis. A single bona fide entry receptor protein supporting this entry mechanism remains elusive. Here we performed proximity ligation of biotin to host cell surface proteins in the vicinity of attached trimeric hemagglutinin-HRP and characterized biotinylated targets using mass spectrometry. This approach identified transferrin receptor 1 (TfR1) as a candidate entry protein. Genetic gain-of-function and loss-of-function experiments, as well as in vitro and in vivo chemical inhibition, confirmed the functional involvement of TfR1 in IAV entry. Recycling deficient mutants of TfR1 do not support entry, indicating that TfR1 recycling is essential for this function. The binding of virions to TfR1 via sialic acids confirmed its role as a directly acting entry factor, but unexpectedly even headless TfR1 promoted IAV particle uptake in trans. TIRF microscopy localized the entering virus-like particles in the vicinity of TfR1. Our data identify TfR1 recycling as a revolving door mechanism exploited by IAV to enter host cells.
Keywords: antiviral; endocytosis; influenza A virus; recycling; transferrin receptor 1.
Conflict of interest statement
The authors declare no competing interest.
Figures


Comment in
-
Could iron deficiency potentiate the appearance of complications in Influenza A virus infection?Proc Natl Acad Sci U S A. 2023 Aug 29;120(35):e2311262120. doi: 10.1073/pnas.2311262120. Epub 2023 Aug 21. Proc Natl Acad Sci U S A. 2023. PMID: 37603764 Free PMC article. No abstract available.
Similar articles
-
Transferrin Receptor Protein 1 Is an Entry Factor for Rabies Virus.J Virol. 2023 Feb 28;97(2):e0161222. doi: 10.1128/jvi.01612-22. Epub 2023 Feb 13. J Virol. 2023. PMID: 36779762 Free PMC article.
-
Transferrin Receptor Protein 1 Cooperates with mGluR2 To Mediate the Internalization of Rabies Virus and SARS-CoV-2.J Virol. 2023 Feb 28;97(2):e0161122. doi: 10.1128/jvi.01611-22. Epub 2023 Feb 13. J Virol. 2023. PMID: 36779763 Free PMC article.
-
Cell binding, uptake, and infection of influenza A virus using recombinant antibody-based receptors.J Virol. 2025 May 20;99(5):e0227524. doi: 10.1128/jvi.02275-24. Epub 2025 Apr 10. J Virol. 2025. PMID: 40207931 Free PMC article.
-
The transferrin receptor: the cellular iron gate.Metallomics. 2017 Oct 18;9(10):1367-1375. doi: 10.1039/c7mt00143f. Metallomics. 2017. PMID: 28671201 Review.
-
Structural analysis of the transferrin receptor multifaceted ligand(s) interface.Biophys Chem. 2019 Nov;254:106242. doi: 10.1016/j.bpc.2019.106242. Epub 2019 Aug 1. Biophys Chem. 2019. PMID: 31419721 Review.
Cited by
-
A live cell imaging-based assay for tracking particle uptake by clathrin-mediated endocytosis.Methods Enzymol. 2024;700:413-454. doi: 10.1016/bs.mie.2024.02.010. Epub 2024 Mar 22. Methods Enzymol. 2024. PMID: 38971609 Free PMC article.
-
Inhibition of early EHDV2-Ibaraki infection steps in bovine cells by endosome alkalinization or ikarugamycin, but not by blockage of individual endocytic pathways.Front Cell Infect Microbiol. 2025 Feb 6;15:1494200. doi: 10.3389/fcimb.2025.1494200. eCollection 2025. Front Cell Infect Microbiol. 2025. PMID: 39981379 Free PMC article.
-
The role of ferroptosis in virus infections.Front Microbiol. 2023 Nov 23;14:1279655. doi: 10.3389/fmicb.2023.1279655. eCollection 2023. Front Microbiol. 2023. PMID: 38075884 Free PMC article. Review.
-
Molecular Evolution of the H5 and H7 Highly Pathogenic Avian Influenza Virus Haemagglutinin Cleavage Site Motif.Rev Med Virol. 2025 Jan;35(1):e70012. doi: 10.1002/rmv.70012. Rev Med Virol. 2025. PMID: 39730318 Free PMC article. Review.
-
Transferrin receptors.Exp Mol Med. 2025 Apr;57(4):724-732. doi: 10.1038/s12276-025-01436-x. Epub 2025 Apr 22. Exp Mol Med. 2025. PMID: 40263550 Free PMC article. Review.
References
-
- Jolly C. L., Sattentau Q. J., Attachment factors. Adv. Exp. Med. Biol. 790, 1–23 (2013). - PubMed
-
- Yamauchi Y., Helenius A., Virus entry at a glance. J. Cell Sci. 126, 1289–1295 (2013). - PubMed
-
- Sempere Borau M., Stertz S., Entry of influenza A virus into host cells–Recent progress and remaining challenges. Curr. Opin. Virol. 48, 23–29 (2021). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

