Plasma proteome perturbation for CMV DNAemia in kidney transplantation
- PMID: 37205661
- PMCID: PMC10198483
- DOI: 10.1371/journal.pone.0285870
Plasma proteome perturbation for CMV DNAemia in kidney transplantation
Abstract
Background: Cytomegalovirus (CMV) infection, either de novo or as reactivation after allotransplantation and chronic immunosuppression, is recognized to cause detrimental alloimmune effects, inclusive of higher susceptibility to graft rejection and substantive impact on chronic graft injury and reduced transplant survival. To obtain further insights into the evolution and pathogenesis of CMV infection in an immunocompromised host we evaluated changes in the circulating host proteome serially, before and after transplantation, and during and after CMV DNA replication (DNAemia), as measured by quantitative polymerase chain reaction (QPCR).
Methods: LC-MS-based proteomics was conducted on 168 serially banked plasma samples, from 62 propensity score-matched kidney transplant recipients. Patients were stratified by CMV replication status into 31 with CMV DNAemia and 31 without CMV DNAemia. Patients had blood samples drawn at protocol times of 3- and 12-months post-transplant. Additionally, blood samples were also drawn before and 1 week and 1 month after detection of CMV DNAemia. Plasma proteins were analyzed using an LCMS 8060 triple quadrupole mass spectrometer. Further, public transcriptomic data on time matched PBMCs samples from the same patients was utilized to evaluate integrative pathways. Data analysis was conducted using R and Limma.
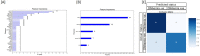
Results: Samples were segregated based on their proteomic profiles with respect to their CMV Dnaemia status. A subset of 17 plasma proteins was observed to predict the onset of CMV at 3 months post-transplant enriching platelet degranulation (FDR, 4.83E-06), acute inflammatory response (FDR, 0.0018), blood coagulation (FDR, 0.0018) pathways. An increase in many immune complex proteins were observed at CMV infection. Prior to DNAemia the plasma proteome showed changes in the anti-inflammatory adipokine vaspin (SERPINA12), copper binding protein ceruloplasmin (CP), complement activation (FDR = 0.03), and proteins enriched in the humoral (FDR = 0.01) and innate immune responses (FDR = 0.01).
Conclusion: Plasma proteomic and transcriptional perturbations impacting humoral and innate immune pathways are observed during CMV infection and provide biomarkers for CMV disease prediction and resolution. Further studies to understand the clinical impact of these pathways can help in the formulation of different types and duration of anti-viral therapies for the management of CMV infection in the immunocompromised host.
Copyright: © 2023 Sigdel et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

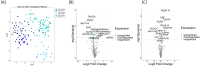


References
-
- Humar A, Snydman D, Practice ASTIDCo. Cytomegalovirus in solid organ transplant recipients. American journal of transplantation: official journal of the American Society of Transplantation and the American Society of Transplant Surgeons 2009;9 Suppl 4:S78–86. - PubMed

