Sequence Variation of Candida albicans Sap2 Enhances Fungal Pathogenicity via Complement Evasion and Macrophage M2-Like Phenotype Induction
- PMID: 37211685
- PMCID: PMC10369283
- DOI: 10.1002/advs.202206713
Sequence Variation of Candida albicans Sap2 Enhances Fungal Pathogenicity via Complement Evasion and Macrophage M2-Like Phenotype Induction
Abstract
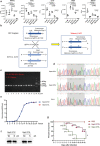
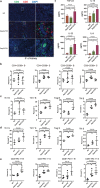
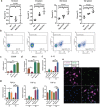
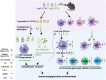
Candida albicans (C. albicans) is an opportunistic pathogen increasingly causing candidiasis worldwide. This study aims to investigate the pattern of systemic immune responses triggered by C. albicans with disease associated variation of Sap2, identifying the novel evasion strategies utilized by clinical isolates. Specifically, a variation in clinical isolates is identified at nucleotide position 817 (G to T). This homozygous variation causes the 273rd amino acid exchange from valine to leucine, close to the proteolytic activation center of Sap2. The mutant (Sap2-273L) generated from SC5314 (Sap2-273V) background carrying the V273L variation within Sap2 displays higher pathogenicity. In comparison to mice infected with Sap2-273V strain, mice infected with Sap2-273L exhibit less complement activation indicated by less serum C3a generation and weaker C3b deposition in the kidney. This inhibitory effect is mainly achieved by Sap2273L -mediated stronger degradation of C3 and C3b. Furthermore, mice infected with Sap2-273L strain exhibit more macrophage phenotype switching from M0 to M2-like and more TGF-β release which further influences T cell responses, generating an immunosuppressed cellular microenvironment characterized by more Tregs and exhausted T cell formation. In summary, the disease-associated sequence variation of Sap2 enhances pathogenicity by complement evasion and M2-like phenotype switching, promoting a more efficient immunosuppressed microenvironment.
Keywords: Candida albicans; complement evasion; fungal pathogenicity; immunosuppressed cellular environment; sequence variation.
© 2023 The Authors. Advanced Science published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- a) Hirakawa M. P., Martinez D. A., Sakthikumar S., Anderson M. Z., Berlin A., Gujja S., Zeng Q., Zisson E., Wang J. M., Greenberg J. M., Berman J., Bennett R. J., Cuomo C. A., Genome Res. 2015, 25, 413; - PMC - PubMed
- b) Zhang C., Wang W., Kong Q., Liu F., Chen J., Sang H., Mycologia 2019, 111, 942; - PubMed
- c) Sanglard D., Ischer F., Calabrese D., Micheli M., Bille J., Drug Resistance Updates 1998, 1, 255. - PubMed
-
- a) Alonso‐Valle H., Acha O., García‐Palomo J. D., Fariñas‐Alvarez C., Fernández‐Mazarrasa C., Fariñas M. C., Eur. J. Clin. Microbiol. Infect. Dis. 2003, 22, 254; - PubMed
- b) Gudlaugsson O., Gillespie S., Lee K., Berg J. V., Hu J., Messer S., Herwaldt L., Pfaller M., Diekema D., Clin. Infect. Dis. 2003, 37, 1172; - PubMed
- c) Pappas P. G., Rex J. H., Lee J., Hamill R. J., Larsen R. A., Powderly W., Kauffman C. A., Hyslop N., Mangino J. E., Chapman S., Horowitz H. W., Edwards J. E., Dismukes W. E., Clin. Infect. Dis. 2003, 37, 634. - PubMed
-
- a) Bacher P., Hohnstein T., Beerbaum E., Röcker M., Blango M. G., Kaufmann S., Röhmel J., Eschenhagen P., Grehn C., Seidel K., Rickerts V., Lozza L., Stervbo U., Nienen M., Babel N., Milleck J., Assenmacher M., Cornely O. A., Ziegler M., Wisplinghoff H., Heine G., Worm M., Siegmund B., Maul J., Creutz P., Tabeling C., Ruwwe‐Glösenkamp C., Sander L. E., Knosalla C., Brunke S., et al., Cell 2019, 176, 1340; - PubMed
- b) Chen J., He R., Sun W., Gao R., Peng Q., Zhu L., Du Y., Ma X., Guo X., Zhang H., Tan C., Wang J., Zhang W., Weng X., Man J., Bauer H., Wang Q. K., Martin B. N., Zhang C. J., Li X., Wang C., Nat. Commun. 2020, 11, 1913. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
