Inhibition of SARS-CoV-2-mediated thromboinflammation by CLEC2.Fc
- PMID: 37211986
- PMCID: PMC10331576
- DOI: 10.15252/emmm.202216351
Inhibition of SARS-CoV-2-mediated thromboinflammation by CLEC2.Fc
Abstract
Thromboinflammation is the major cause of morbidity and mortality in COVID-19 patients, and post-mortem examination demonstrates the presence of platelet-rich thrombi and microangiopathy in visceral organs. Moreover, persistent microclots were detected in both acute COVID-19 and long COVID plasma samples. However, the molecular mechanism of SARS-CoV-2-induced thromboinflammation is still unclear. We found that the spleen tyrosine kinase (Syk)-coupled C-type lectin member 2 (CLEC2), which was highly expressed in platelets and alveolar macrophages, interacted with the receptor-binding domain (RBD) of SARS-CoV-2 spike protein (SARS-CoV-2 RBD) directly. Unlike the thread-like NETs, SARS-CoV-2-induced aggregated NET formation in the presence of wild-type (WT), but not CLEC2-deficient platelets. Furthermore, SARS-CoV-2 spike pseudotyped lentivirus was able to induce NET formation via CLEC2, indicating SARS-CoV-2 RBD engaged CLEC2 to activate platelets to enhance NET formation. Administration of CLEC2.Fc inhibited SARS-CoV-2-induced NET formation and thromboinflammation in AAV-ACE2-infected mice. Thus, CLEC2 is a novel pattern recognition receptor for SARS-CoV-2, and CLEC2.Fc and may become a promising therapeutic agent to inhibit SARS-CoV-2-induced thromboinflammation and reduced the risk of post-acute sequelae of COVID-19 (PASC) in the future.
Keywords: CLEC2; NET; SARS-CoV-2; platelet; thromboinflammation.
© 2023 The Authors. Published under the terms of the CC BY 4.0 license.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
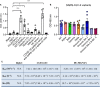
- A, B
Interaction between several human C‐type lectins (50 μg/ml) and recombinant SARS‐CoV‐2 RBD (50 μl/well) was determined by ELISA (A), and the interactions of hCLEC2.Fc with SARS‐CoV‐2 RBDs from various strains (B).
- C
The binding affinity of SARS‐CoV‐2‐spike RBD and human CLEC2.Fc fusion protein was measured by biolayer interferometry. Ka: Rate constants for association. Kd: Rate constants for dissociation. KD: Equilibrium dissociation constant.

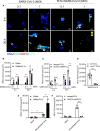
- A
The image of NETs (arrowheads: aggregated NETs).
- B
Quantitation of NET area by citrullinated Histone H3 (Cit‐H3) area.
- C
Human platelets were pretreated with anti‐CLEC2 mAb (AYP1, 2.5 μg/ml) or isotype control for 15 min at room temperature before incubation with neutrophils and SARS‐CoV‐2 stimulation.
- D
Neutrophils (4 × 105/ml) from wild type (WT) mice were incubated with SARS‐CoV‐2 (MOI = 1) in the presence of WT platelets (4 × 106/ml) or clec2 −/− platelets (4 × 106/ml) for 5 h at 37°C.
- E, F
Human neutrophils were stimulated with SARS‐CoV‐2 spike RBD pseudotyped virus (pseudotyped virus; MOI = 0.1) in presence of autologous platelets at 37°C for 3 h (E). Platelets were incubated with anti‐CLEC2 mAb (2.5 μg/ml) at room temperature for 15 min before incubation with neutrophils and pseudotyped virus stimulation at 37°C for 3 h (F). The level of NET formation was determined using Cit‐H3 area (μm2).

- A, B
Human neutrophils were stimulated with SARS‐CoV‐2 (MOI = 0.1 and 1) at 37°C in the presence of isotype control (hIgG1, 3 μg/ml) or human CLEC2.Fc (3 μg/ml) and with or without autologous platelets for 5 h (A) or with or without anti‐SARS‐CoV‐2 spike RBD antibody (3 μg/ml) for 20 h (B). Data are means ± SD from at least three independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 (Student's t‐test).
- C–E
C57BL/6 mice were inoculated with AAV‐hACE2 at day 14 before the SARS‐CoV‐2 challenge. For the prophylactic use of hCLEC2.Fc, mice were intravenously (i.v.) injected with CLEC2.Fc (100 μg/mouse) then intratracheally injected with SARS‐CoV‐2 (8 × 104 PFU). For the therapeutic use of hCLEC2.Fc, mice were intratracheally inoculated with SARS‐CoV‐2 (8 × 104 PFU) and intraperitoneally injected with CLEC2.Fc (100 μg/per mice). Lung tissues were collected at day 3 day and day 5 post‐infection. (C) The expression levels of viral copy number (D), cytokines and chemokines in lung were measured by qPCR (E).
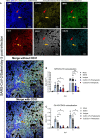
- A
AAV‐hACE2 mice were treated with saline and challenged with SARS‐CoV‐2 (8 × 104 PFU) for 5 days. Lung tissue sections were stained with DAPI (blue), anti‐MPO antibody (green), anti‐citrullinated histone H3 (Cit‐H3) antibody (red), anti‐CD42b antibody (yellow) and anti‐CD31 antibody (gray). yellow arrow: NETs (DNA+Gr‐1+MPO+Cit‐H3+); red arrow: immunothrombosis (NETs + thrombus (CD42b+)), Scale bar is 100 μm.
- B, C
For prophylactic treatment, CLEC2.Fc (or vehicle) was given at 1 h before virus challenge; for therapeutic treatment, CLEC2.Fc was given at 8 h post‐infection. The area of NET (colocalization area of MPO and Cit‐H3) (B) and immunothrombosis (colocalization area of Cit‐H3 and CD42b) (C) were measured using MetaMorph software.
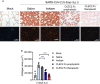
- A, B
AAV‐hACE2 mice were treated with saline and challenged with SARS‐CoV‐2 (8 × 104 PFU) for 5 days. Samples were collected at day 5 post‐infection, and the collagen was stained with Picro Sirius red (A), while collagen deposition was quantified using MetaMorph software and presented as area (μm2) (B). The images were captured by light microscopy (upper panel) and polarized light microscopy (lower panel).
Similar articles
-
CLEC5A and TLR2 are critical in SARS-CoV-2-induced NET formation and lung inflammation.J Biomed Sci. 2022 Jul 11;29(1):52. doi: 10.1186/s12929-022-00832-z. J Biomed Sci. 2022. PMID: 35820906 Free PMC article.
-
What Role Does Microthrombosis Play in Long COVID?Semin Thromb Hemost. 2024 Jun;50(4):527-536. doi: 10.1055/s-0043-1774795. Epub 2023 Sep 25. Semin Thromb Hemost. 2024. PMID: 37748518 Review.
-
C-type lectins and extracellular vesicles in virus-induced NETosis.J Biomed Sci. 2021 Jun 11;28(1):46. doi: 10.1186/s12929-021-00741-7. J Biomed Sci. 2021. PMID: 34116654 Free PMC article. Review.
-
Thromboinflammation in long COVID-the elusive key to postinfection sequelae?J Thromb Haemost. 2023 Aug;21(8):2020-2031. doi: 10.1016/j.jtha.2023.04.039. Epub 2023 May 11. J Thromb Haemost. 2023. PMID: 37178769 Free PMC article. Review.
-
Extracellular vesicles from CLEC2-activated platelets enhance dengue virus-induced lethality via CLEC5A/TLR2.Nat Commun. 2019 Jun 3;10(1):2402. doi: 10.1038/s41467-019-10360-4. Nat Commun. 2019. PMID: 31160588 Free PMC article.
Cited by
-
SARS-CoV-2 Omicron variant infection affects blood platelets, a comparative analysis with Delta variant.Front Immunol. 2023 Sep 27;14:1231576. doi: 10.3389/fimmu.2023.1231576. eCollection 2023. Front Immunol. 2023. PMID: 37828997 Free PMC article.
-
Neutrophil extracellular trap formation during surgical procedures: a pilot study.Sci Rep. 2023 Sep 14;13(1):15217. doi: 10.1038/s41598-023-42565-5. Sci Rep. 2023. PMID: 37709941 Free PMC article.
-
COVID-19: a vascular nightmare unfolding.Front Immunol. 2025 Aug 1;16:1593885. doi: 10.3389/fimmu.2025.1593885. eCollection 2025. Front Immunol. 2025. PMID: 40821797 Free PMC article. Review.
References
-
- Andrews RK, Arthur JF, Gardiner E (2014) Neutrophil extracellular traps (NETs) and the role of platelets in infection. Thromb Haemost 112: 659–665 - PubMed
-
- Chen ST, Lin YL, Huang MT, Wu MF, Cheng SC, Lei HY, Lee CK, Chiou TW, Wong CH, Hsieh SL (2008) CLEC5A is critical for dengue‐virus‐induced lethal disease. Nature 453: 672–676 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

