Integrative analysis of gene expression and DNA methylation to identify biomarkers of non-genital warts induced by low-risk human papillomaviruses infection
- PMID: 37215908
- PMCID: PMC10196596
- DOI: 10.1016/j.heliyon.2023.e16101
Integrative analysis of gene expression and DNA methylation to identify biomarkers of non-genital warts induced by low-risk human papillomaviruses infection
Abstract
Background: Human papillomaviruses have been shown to dysregulate the gene expression and DNA methylation profiles of their host cells over the course of infection. However, there is a lack of information on the impact of low-risk HPV infection and wart formation on host cell's expression and methylation patterns. Therefore, the objective of this study is to analyse the genome and methylome of common warts using an integrative approach.
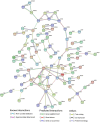
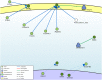
Methods: In the present study, gene expression (GSE136347) and methylation (GSE213888) datasets of common warts were obtained from the GEO database. Identification of the differentially expressed and differentially methylated genes was carried out using the RnBeads R package and the edgeR Bioconductor package. Next, functional annotation of the identified genes was obtained using the Database for Annotation, Visualization, and Integrated Discovery (DAVID). Network construction and analyses of the gene-gene, protein-protein, and signaling interactions of the differentially expressed and differentially methylated genes was performed using the GeneMANIA web interface, the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) database, and the Signaling Network Open Resource 2.0 (SIGNOR 2.0), respectively. Lastly, significant hub genes were identified using the Cytoscape application CytoHubba.
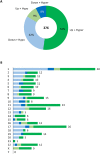
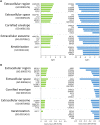
Results: A total of 276 genes were identified as differentially expressed and differentially methylated in common warts, with 52% being upregulated and hypermethylated. Functional enrichment analysis identified extracellular components as the most enriched annotations, while network analyses identified ELN, ITGB1, TIMP1, MMP2, LGALS3, COL1A1 and ANPEP as significant hub genes.
Conclusions: To the best knowledge of the authors, this is the first integrative study to be carried out on non-genital warts induced by low-risk HPV types. Future studies are required to re-validate the findings in larger populations using alternative approaches.
Keywords: DNA methylation; Gene expression; HPV; Human papillomavirus; Warts.
© 2023 The Authors. Published by Elsevier Ltd.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


Similar articles
-
Identification of abnormally methylated-differentially expressed genes and pathways in osteoarthritis: a comprehensive bioinformatic study.Clin Rheumatol. 2021 Aug;40(8):3247-3256. doi: 10.1007/s10067-020-05539-w. Epub 2021 Jan 9. Clin Rheumatol. 2021. PMID: 33420869
-
Genome-Wide CpG Island Methylation Profiles of Cutaneous Skin with and without HPV Infection.Int J Mol Sci. 2019 Sep 28;20(19):4822. doi: 10.3390/ijms20194822. Int J Mol Sci. 2019. PMID: 31569353 Free PMC article.
-
Identification of key DNA methylation-driven genes in prostate adenocarcinoma: an integrative analysis of TCGA methylation data.J Transl Med. 2019 Sep 18;17(1):311. doi: 10.1186/s12967-019-2065-2. J Transl Med. 2019. PMID: 31533842 Free PMC article.
-
Genome-Wide Tiling Array Analysis of HPV-Induced Warts Reveals Aberrant Methylation of Protein-Coding and Non-Coding Regions.Genes (Basel). 2019 Dec 27;11(1):34. doi: 10.3390/genes11010034. Genes (Basel). 2019. PMID: 31892232 Free PMC article.
-
Common gene signatures and key pathways in hypopharyngeal and esophageal squamous cell carcinoma: Evidence from bioinformatic analysis.Medicine (Baltimore). 2020 Oct 16;99(42):e22434. doi: 10.1097/MD.0000000000022434. Medicine (Baltimore). 2020. PMID: 33080677 Free PMC article.
References
-
- Luria L., Cardoza-Favarato G. StatPearls. StatPearls Publishing; Treasure Island (FL: 2022. Human papillomavirus.http://www.ncbi.nlm.nih.gov/books/NBK448132/ accessed June 2, 2022)
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

