Origin and segregation of the human germline
- PMID: 37217306
- PMCID: PMC10203729
- DOI: 10.26508/lsa.202201706
Origin and segregation of the human germline
Abstract
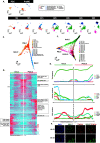
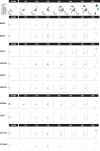
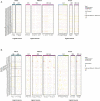
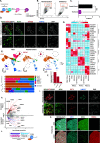
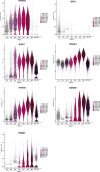
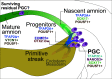
Human germline-soma segregation occurs during weeks 2-3 in gastrulating embryos. Although direct studies are hindered, here, we investigate the dynamics of human primordial germ cell (PGCs) specification using in vitro models with temporally resolved single-cell transcriptomics and in-depth characterisation using in vivo datasets from human and nonhuman primates, including a 3D marmoset reference atlas. We elucidate the molecular signature for the transient gain of competence for germ cell fate during peri-implantation epiblast development. Furthermore, we show that both the PGCs and amnion arise from transcriptionally similar TFAP2A-positive progenitors at the posterior end of the embryo. Notably, genetic loss of function experiments shows that TFAP2A is crucial for initiating the PGC fate without detectably affecting the amnion and is subsequently replaced by TFAP2C as an essential component of the genetic network for PGC fate. Accordingly, amniotic cells continue to emerge from the progenitors in the posterior epiblast, but importantly, this is also a source of nascent PGCs.
© 2023 Castillo-Venzor et al.
Conflict of interest statement
WWC Tang is currently employed by Adrestia Therapeutics Ltd. The other authors declare that they have no conflict of interest.
Figures









Similar articles
-
On the origin of the human germline.Development. 2018 Jul 23;145(16):dev150433. doi: 10.1242/dev.150433. Development. 2018. PMID: 30037844 Review.
-
The Germ Cell Fate of Cynomolgus Monkeys Is Specified in the Nascent Amnion.Dev Cell. 2016 Oct 24;39(2):169-185. doi: 10.1016/j.devcel.2016.09.007. Epub 2016 Oct 6. Dev Cell. 2016. PMID: 27720607
-
Does mouse embryo primordial germ cell activation start before implantation as suggested by single-cell transcriptomics dynamics?Mol Hum Reprod. 2016 Mar;22(3):208-25. doi: 10.1093/molehr/gav072. Epub 2016 Jan 5. Mol Hum Reprod. 2016. PMID: 26740066
-
Human blastocyst outgrowths recapitulate primordial germ cell specification events.Mol Hum Reprod. 2019 Sep 1;25(9):519-526. doi: 10.1093/molehr/gaz035. Mol Hum Reprod. 2019. PMID: 31211841 Free PMC article.
-
Primordial germ cell specification: a context-dependent cellular differentiation event [corrected].Philos Trans R Soc Lond B Biol Sci. 2014 Dec 5;369(1657):20130543. doi: 10.1098/rstb.2013.0543. Philos Trans R Soc Lond B Biol Sci. 2014. PMID: 25349452 Free PMC article. Review.
Cited by
-
The emergence of human primordial germ cell-like cells in stem cell-derived gastruloids.Sci Adv. 2025 Mar 28;11(13):eado1350. doi: 10.1126/sciadv.ado1350. Epub 2025 Mar 26. Sci Adv. 2025. PMID: 40138398 Free PMC article.
-
Early human development and stem cell-based human embryo models.Cell Stem Cell. 2024 Oct 3;31(10):1398-1418. doi: 10.1016/j.stem.2024.09.002. Cell Stem Cell. 2024. PMID: 39366361 Free PMC article. Review.
-
X-Linked Gene Dosage and SOX2 Act as Key Roadblocks for Human Germ Cell Specification in Klinefelter Syndrome.Adv Sci (Weinh). 2025 Apr;12(15):e2410533. doi: 10.1002/advs.202410533. Epub 2025 Feb 25. Adv Sci (Weinh). 2025. PMID: 39996497 Free PMC article.
-
Genetic and genomic insights into male reproductive tract development.Fertil Steril. 2025 Jun;123(6):970-979. doi: 10.1016/j.fertnstert.2025.03.024. Epub 2025 Mar 31. Fertil Steril. 2025. PMID: 40174856
-
Efficient derivation of embryonic stem cells and primordial germ cell-like cells in cattle.J Reprod Dev. 2024 Apr 4;70(2):82-95. doi: 10.1262/jrd.2023-087. Epub 2024 Feb 15. J Reprod Dev. 2024. PMID: 38355134 Free PMC article.
References
-
- Bryja J, Konečný A (2003) Fast sex identification in wild mammals using PCR amplification of the Sry gene. Folia Zool Brno 52: 269–274.
-
- Bullen P, Wilson DI, Strachan T, Lindsay S, Wilson DI (1997) Molecular genetics of early human development The Carnegie Staging of Human Embryos: A Practical Guide. Bios: 265.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
