Structural genomics of the human dopamine receptor system
- PMID: 37221270
- PMCID: PMC10397222
- DOI: 10.1038/s41422-023-00808-0
Structural genomics of the human dopamine receptor system
Abstract
The dopaminergic system, including five dopamine receptors (D1R to D5R), plays essential roles in the central nervous system (CNS); and ligands that activate dopamine receptors have been used to treat many neuropsychiatric disorders, including Parkinson's Disease (PD) and schizophrenia. Here, we report cryo-EM structures of all five subtypes of human dopamine receptors in complex with G protein and bound to the pan-agonist, rotigotine, which is used to treat PD and restless legs syndrome. The structures reveal the basis of rotigotine recognition in different dopamine receptors. Structural analysis together with functional assays illuminate determinants of ligand polypharmacology and selectivity. The structures also uncover the mechanisms of dopamine receptor activation, unique structural features among the five receptor subtypes, and the basis of G protein coupling specificity. Our work provides a comprehensive set of structural templates for the rational design of specific ligands to treat CNS diseases targeting the dopaminergic system.
© 2023. The Author(s) under exclusive licence to Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences.
Conflict of interest statement
The authors declare no competing interests.
Figures




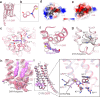


References
-
- Robbins TW. Dopamine and cognition. Curr. Opin. Neurol. 2003;16:S1–S2. - PubMed
-
- Volkow ND, et al. Association between decline in brain dopamine activity with age and cognitive and motor impairment in healthy individuals. Am. J. Psychiatry. 1998;155:344–349. - PubMed
-
- Koob GF. Dopamine, addiction and reward. Semin. Neurosci. 1992;4:139–148.
-
- Neve, K. A. & Neve, R. L. Molecular biology of dopamine receptors. In: Neve, K. A. & Neve, R. L. (eds) The Dopamine Receptors. The Receptors. (Humana Press, Totowa, 1997).
-
- Bonuccelli U, Del Dotto P, Rascol O. Role of dopamine receptor agonists in the treatment of early Parkinson’s disease. Parkinsonism Relat. Disord. 2009;15:S44–S53. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

