SVcnn: an accurate deep learning-based method for detecting structural variation based on long-read data
- PMID: 37221476
- PMCID: PMC10207598
- DOI: 10.1186/s12859-023-05324-x
SVcnn: an accurate deep learning-based method for detecting structural variation based on long-read data
Abstract
Background: Structural variations (SVs) refer to variations in an organism's chromosome structure that exceed a length of 50 base pairs. They play a significant role in genetic diseases and evolutionary mechanisms. While long-read sequencing technology has led to the development of numerous SV caller methods, their performance results have been suboptimal. Researchers have observed that current SV callers often miss true SVs and generate many false SVs, especially in repetitive regions and areas with multi-allelic SVs. These errors are due to the messy alignments of long-read data, which are affected by their high error rate. Therefore, there is a need for a more accurate SV caller method.
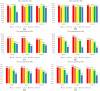
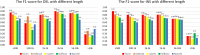
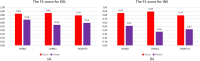
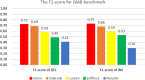
Result: We propose a new method-SVcnn, a more accurate deep learning-based method for detecting SVs by using long-read sequencing data. We run SVcnn and other SV callers in three real datasets and find that SVcnn improves the F1-score by 2-8% compared with the second-best method when the read depth is greater than 5×. More importantly, SVcnn has better performance for detecting multi-allelic SVs.
Conclusions: SVcnn is an accurate deep learning-based method to detect SVs. The program is available at https://github.com/nwpuzhengyan/SVcnn .
Keywords: Deep learning; Long-read sequencing data; SV caller; Structural variations.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


Similar articles
-
FindCSV: a long-read based method for detecting complex structural variations.BMC Bioinformatics. 2024 Sep 28;25(1):315. doi: 10.1186/s12859-024-05937-w. BMC Bioinformatics. 2024. PMID: 39342151 Free PMC article.
-
SVsearcher: A more accurate structural variation detection method in long read data.Comput Biol Med. 2023 May;158:106843. doi: 10.1016/j.compbiomed.2023.106843. Epub 2023 Mar 31. Comput Biol Med. 2023. PMID: 37019014
-
SVvalidation: A long-read-based validation method for genomic structural variation.PLoS One. 2024 Jan 5;19(1):e0291741. doi: 10.1371/journal.pone.0291741. eCollection 2024. PLoS One. 2024. PMID: 38181020 Free PMC article.
-
Detection of somatic structural variants from short-read next-generation sequencing data.Brief Bioinform. 2021 May 20;22(3):bbaa056. doi: 10.1093/bib/bbaa056. Brief Bioinform. 2021. PMID: 32379294 Free PMC article. Review.
-
Structural variation detection using next-generation sequencing data: A comparative technical review.Methods. 2016 Jun 1;102:36-49. doi: 10.1016/j.ymeth.2016.01.020. Epub 2016 Feb 1. Methods. 2016. PMID: 26845461 Review.
Cited by
-
Unlocking precision medicine: clinical applications of integrating health records, genetics, and immunology through artificial intelligence.J Biomed Sci. 2025 Feb 7;32(1):16. doi: 10.1186/s12929-024-01110-w. J Biomed Sci. 2025. PMID: 39915780 Free PMC article. Review.
-
invMap: a sensitive mapping tool for long noisy reads with inversion structural variants.Bioinformatics. 2023 Dec 1;39(12):btad726. doi: 10.1093/bioinformatics/btad726. Bioinformatics. 2023. PMID: 38058196 Free PMC article.
-
Comparative population pangenomes reveal unexpected complexity and fitness effects of structural variants.bioRxiv [Preprint]. 2025 Feb 13:2025.02.11.637762. doi: 10.1101/2025.02.11.637762. bioRxiv. 2025. PMID: 39990470 Free PMC article. Preprint.
-
FindCSV: a long-read based method for detecting complex structural variations.BMC Bioinformatics. 2024 Sep 28;25(1):315. doi: 10.1186/s12859-024-05937-w. BMC Bioinformatics. 2024. PMID: 39342151 Free PMC article.
-
LcDel: deletion variation detection based on clustering and long reads.Front Genet. 2024 May 10;15:1404415. doi: 10.3389/fgene.2024.1404415. eCollection 2024. Front Genet. 2024. PMID: 38798694 Free PMC article.
References
-
- Rovelet-Lecrux A, Hannequin D, Raux G, Meur NL, Laquerrière A, Vital A, Dumanchin C, Feuillette S, Brice A, Vercelletto M. App locus duplication causes autosomal dominant early-onset alzheimer disease with cerebral amyloid angiopathy. Nature Genet. 2006;38(1):24–26. doi: 10.1038/ng1718. - DOI - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources

