Nonbacterial Microflora in Wastewater Treatment Plants: an Underappreciated Potential Source of Pathogens
- PMID: 37222623
- PMCID: PMC10269893
- DOI: 10.1128/spectrum.00481-23
Nonbacterial Microflora in Wastewater Treatment Plants: an Underappreciated Potential Source of Pathogens
Abstract
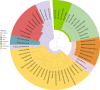
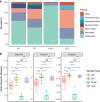
Wastewater treatment plants (WWTPs) receive and treat large volumes of domestic, industrial, and urban wastewater containing pathogenic and nonpathogenic microorganisms, chemical compounds, heavy metals, and other potentially hazardous substances. WWTPs play an essential role in preserving human, animal, and environmental health by removing many of these toxic and infectious agents, particularly biological hazards. Wastewater contains complex consortiums of bacterial, viral, archaeal, and eukaryotic species, and while bacteria in WWTP have been extensively studied, the temporal and spatial distribution of nonbacterial microflora (viruses, archaea, and eukaryotes) is less understood. In this study, we analyzed the viral, archaeal, and eukaryotic microflora in wastewater throughout a treatment plant (raw influent, effluent, oxidation pond water, and oxidation pond sediment) in Aotearoa (New Zealand) using Illumina shotgun metagenomic sequencing. Our results suggest a similar trend across many taxa, with an increase in relative abundance in oxidation pond samples compared to influent and effluent samples, except for archaea, which had the opposite trend. Additionally, some microbial families, such as Podoviridae bacteriophages and Apicomplexa alveolates, appeared largely unaffected by the treatment process, with their relative abundance remaining stable throughout. Several groups encompassing pathogenic species, such as Leishmania, Plasmodium, Toxoplasma, Apicomplexa, Cryptococcus, Botrytis, and Ustilago, were identified. If present, these potentially pathogenic species could be a threat to human and animal health and agricultural productivity; therefore, further investigation is warranted. These nonbacterial pathogens should be considered when assessing the potential for vector transmission, distribution of biosolids to land, and discharge of treated wastewater to waterways or land. IMPORTANCE Nonbacterial microflora in wastewater remain understudied compared to their bacterial counterparts despite their importance in the wastewater treatment process. In this study, we report the temporal and spatial distributions of DNA viruses, archaea, protozoa, and fungi in raw wastewater influent, effluent, oxidation pond water, and oxidation pond sediments by using shotgun metagenomic sequencing. Our study indicated the presence of groups of nonbacterial taxa which encompass pathogenic species that may have potential to cause disease in humans, animals, and agricultural crops. We also observed higher alpha diversity in viruses, archaea, and fungi in effluent samples than in influent samples. This suggests that the resident microflora in the wastewater treatment plant may be making a greater contribution to the diversity of taxa observed in wastewater effluent than previously thought. This study provides important insights to better understand the potential human, animal, and environmental health impacts of discharged treated wastewater.
Keywords: One Health; archaea; environmental and human health; fungi; nonbacterial microflora; protozoa; shotgun metagenomic sequencing; virus; wastewater treatment plant.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Gude VG. 2015. A new perspective on microbiome and resource management in wastewater systems. Biotechnol Biomater 5:184. doi: 10.4172/2155-952X.1000184. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

