Zinc finger and SCAN domain-containing protein 18 is a potential DNA methylation-modified tumor suppressor and biomarker in breast cancer
- PMID: 37223020
- PMCID: PMC10200902
- DOI: 10.3389/fendo.2023.1095604
Zinc finger and SCAN domain-containing protein 18 is a potential DNA methylation-modified tumor suppressor and biomarker in breast cancer
Abstract
Introduction: Zinc finger and SCAN domain-containing protein 18 (ZSCAN18) has been investigated as a putative biomarker of multiple human cancers. However, the expression profile, epigenetic modification, prognostic value, transcription regulation, and molecular mechanism of ZSCAN18 in breast cancer (BC) remain unknown.
Methods: In the study, we present an integrated analysis of ZSCAN18 in BC based on public omics datasets with the use of multiple bioinformatics tools. Genes potentially regulated through restoration of ZSCAN18 expression in MDA-MB-231 cells were investigated to identify pathways associated with BC.
Results: We observed that ZSCAN18 was downregulated in BC and mRNA expression was significantly correlated with clinicopathological parameters. Low expression of ZSCAN18 was found in the HER2-positive and TNBC subtypes. High expression of ZSCAN18 was associated with good prognosis. As compared to normal tissues, the extent of ZSCAN18 DNA methylation was greater with fewer genetic alterations in BC tissues. ZSCAN18 was identified as a transcription factor that might be involved in intracellular molecular and metabolic processes. Low ZSCAN18 expression was associated with the cell cycle and glycolysis signaling pathway. Overexpression of ZSCAN18 inhibited mRNA expression of genes associated with the Wnt/β-catenin and glycolysis signaling pathways, including CTNNB1, BCL9, TSC1, and PFKP. ZSCAN18 expression was negatively correlated with infiltrating B cells and dendritic cells (DCs), as determined by the TIMER web server and reference to the TISIDB. ZSCAN18 DNA methylation was positively correlated with activated B cells, activated CD8+ and CD4+ T cells, macrophages, neutrophils, and activated DCs. Moreover, five ZSCAN18-related hub genes (KDM6B, KAT6A, KMT2D, KDM1A, and HSPBP1) were identified. ZSCAN18, ZNF396, and PGBD1 were identified as components of a physical complex.
Conclusion: ZSCAN18 is a potential tumor suppressor in BC, as expression is modified by DNA methylation and associated with patient survival. In addition, ZSCAN18 plays important roles in transcription regulation, the glycolysis signaling pathway, and the tumor immune microenvironment.
Keywords: DNA methylation; ZSCAN18; breast cancer; integrated analysis; transcription regulation; tumor suppressor.
Copyright © 2023 Wang, Luo, Fu, He, Pan, Fan, Wen and Fan.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


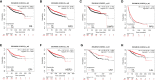




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

