Antimicrobial resistance and virulence genes of invasive Salmonella enterica from children with bacteremia in north-central Nigeria
- PMID: 37223673
- PMCID: PMC10201152
- DOI: 10.1177/20503121231175322
Antimicrobial resistance and virulence genes of invasive Salmonella enterica from children with bacteremia in north-central Nigeria
Abstract
Objectives: Bacteremia due to invasive Salmonella enterica has been reported earlier in children in Nigeria. This study aimed to detect the virulence and antibiotic resistance genes of invasive Salmonella enterica from children with bacteremia in north-central Nigeria.
Method: From June 2015 to June 2018, 4163 blood cultures yielded 83 Salmonella isolates. This is a secondary cross-sectional analysis of the Salmonella isolates. The Salmonella enterica were isolated and identified using standard bacteriology protocol. Biochemical identifications of the Salmonella enterica were made by Phoenix MD 50 identification system. Further identification and confirmation were done with polyvalent antisera O and invA gene. Antimicrobial susceptibility testing was done following clinical and laboratory standard institute guidelines. Resistant genes and virulence genes were determined using a real-time polymerase chain reaction.
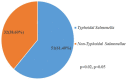
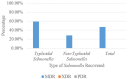
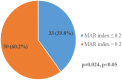
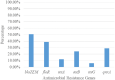
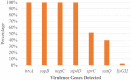
Result: Salmonella typhi 51 (61.4%) was the most prevalent serovar, followed by Salmonella species 13 (15.7%), choleraesuis 8 (9.6%), enteritidis 6 (7.2%), and typhimurium 5 (6.1%). Fifty-one (61.4%) of 83 Salmonella enterica were typhoidal, while 32 (38.6%) were not. Sixty-five (78.3%) of the 83 Salmonella enterica isolates were resistant to ampicillin and trimethoprim-sulfamethoxazole, followed by chloramphenicol 39 (46.7%), tetracycline 41 (41.4%), piperacillin 33 (33.9%), amoxicillin-clavulanate, and streptomycin 21 (25.3%), while cephalothin was 19 (22.9%). Thirty-nine (46.9%) of the 83 Salmonella enterica isolates were multi-drug resistant, and none were extensive drug resistant or pan-drug resistant. A blaTEM 42 (50.6%), floR 32 (38.6%), qnrA 24 (28.9%), tetB 20 (20.1%), tetA 10 (10.0%), and tetG 5 (6.0%) were the antibiotic resistance genes detected. There were perfect agreement between phenotypic and genotypic detection of antimicrobial resistance in tetracycline, ciprofloxacin, and chloramphenicol, while beta-lactam showed κ = 0.60 agreement. All of the Salmonella enterica isolates had the virulence genes invA, sopB, mgtC, and sip4D, while 33 (39.8%), 45 (51.8%), and 2 (2.4%) had ssaQ, spvC, and ljsGI-1, respectively.
Conclusion: Our findings showed multi-drug resistant Salmonella enterica in children with bacteremia in northern Nigeria. In addition, significant virulence and antimicrobial resistance genes were found in invasive Salmonella enterica in northern Nigeria. Thus, our study emphasizes the need to monitor antimicrobial resistance in Salmonella enterica from invasive sources in Nigeria and supports antibiotic prudence.
Keywords: Salmonella typhi; antibiotic resistance; bacteremia; non-typhoidal Salmonella; northern Nigeria.
© The Author(s) 2023.
Conflict of interest statement
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Figures
References
-
- CDC. Serotypes and the Importance of Serotyping Salmonella | Salmonella Atlas | Reports and Publications | Salmonella | CDC. Center for Disease Control and Prevention. Epub ahead of print 2015. DOI: 10.1208/s12249-010-9573-y. - DOI
-
- Igomu EE. Salmonella Kentucky in Nigeria and the Africa continent. African J Clin Exp Microbiol 2020; 21: 272–283.
LinkOut - more resources
Full Text Sources

