The role of site-2-proteases in bacteria: a review on physiology, virulence, and therapeutic potential
- PMID: 37223736
- PMCID: PMC10202637
- DOI: 10.1093/femsml/uqad025
The role of site-2-proteases in bacteria: a review on physiology, virulence, and therapeutic potential
Abstract
Site-2-proteases are a class of intramembrane proteases involved in regulated intramembrane proteolysis. Regulated intramembrane proteolysis is a highly conserved signaling mechanism that commonly involves sequential digestion of an anti-sigma factor by a site-1- and site-2-protease in response to external stimuli, resulting in an adaptive transcriptional response. Variation of this signaling cascade continues to emerge as the role of site-2-proteases in bacteria continues to be explored. Site-2-proteases are highly conserved among bacteria and play a key role in multiple processes, including iron uptake, stress response, and pheromone production. Additionally, an increasing number of site-2-proteases have been found to play a pivotal role in the virulence properties of multiple human pathogens, such as alginate production in Pseudomonas aeruginosa, toxin production in Vibrio cholerae, resistance to lysozyme in enterococci and antimicrobials in several Bacillus spp, and cell-envelope lipid composition in Mycobacterium tuberculosis. The prominent role of site-2-proteases in bacterial pathogenicity highlights the potential of site-2-proteases as novel targets for therapeutic intervention. In this review, we summarize the role of site-2-proteases in bacterial physiology and virulence, as well as evaluate the therapeutic potential of site-2-proteases.
© The Author(s) 2023. Published by Oxford University Press on behalf of FEMS.
Conflict of interest statement
None declared.
Figures

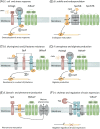
References
-
- Akiyama Y, Kanehara K, Ito K. RseP (YaeL), an Escherichia coli RIP protease, cleaves transmembrane sequences. EMBO J. 2004;23:4434–42. https://doi.org/doi: 10.1038/sj.emboj.7600449. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous
