Integration analysis of miRNA-mRNA expression exploring their potential roles in intrahepatic cholangiocarcinoma
- PMID: 37225858
- PMCID: PMC10209200
- DOI: 10.1038/s41598-023-35288-0
Integration analysis of miRNA-mRNA expression exploring their potential roles in intrahepatic cholangiocarcinoma
Abstract
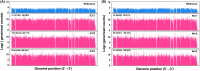
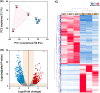
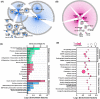
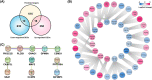
Intrahepatic cholangiocarcinoma (ICC) is the second common primary hepatic malignancy tumor. In this study, an integrative analysis of differentially expressed genes (DEGs) and miRNAs from the ICC onset and adjacent normal tissues were performed to explore the regulatory roles of miRNA-mRNA interaction. A total of 1018 DEGs and 39 miRNAs were likely involved in ICC pathogenesis, suggesting the changes in cell metabolism in ICC development. The built network indicated that 30 DEGs were regulated by 16 differentially expressed miRNA. The screened DEGs and miRNA together were probably considered the biomarkers of ICC, and their important roles in ICC pathogenesis remain to be elucidated. This study could provide a good basis to uncover the regulatory mechanism of miRNA and mRNAs in ICC pathogenesis.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Identification of biomarkers of intrahepatic cholangiocarcinoma via integrated analysis of mRNA and miRNA microarray data.Mol Med Rep. 2017 Mar;15(3):1051-1056. doi: 10.3892/mmr.2017.6123. Epub 2017 Jan 16. Mol Med Rep. 2017. PMID: 28098904 Free PMC article.
-
The role of microRNAs in intrahepatic cholangiocarcinoma.J Cell Mol Med. 2017 Jan;21(1):177-184. doi: 10.1111/jcmm.12951. Epub 2016 Sep 13. J Cell Mol Med. 2017. PMID: 27619971 Free PMC article. Review.
-
CeRNA regulatory network-based analysis to study the roles of noncoding RNAs in the pathogenesis of intrahepatic cholangiocellular carcinoma.Aging (Albany NY). 2020 Jan 17;12(2):1047-1086. doi: 10.18632/aging.102634. Epub 2020 Jan 17. Aging (Albany NY). 2020. PMID: 31956102 Free PMC article.
-
Identification of a novel microRNA signature associated with intrahepatic cholangiocarcinoma (ICC) patient prognosis.BMC Cancer. 2015 Feb 18;15:64. doi: 10.1186/s12885-015-1067-6. BMC Cancer. 2015. PMID: 25880914 Free PMC article.
-
MicroRNAs in the biology and diagnosis of cholangiocarcinoma.Semin Liver Dis. 2015 Feb;35(1):55-62. doi: 10.1055/s-0034-1397349. Epub 2015 Jan 29. Semin Liver Dis. 2015. PMID: 25632935 Review.
Cited by
-
Regulation of lipid metabolism by APOE4 in intrahepatic cholangiocarcinoma via the enhancement of ABCA1 membrane expression.PeerJ. 2024 Jan 22;12:e16740. doi: 10.7717/peerj.16740. eCollection 2024. PeerJ. 2024. PMID: 38274331 Free PMC article.
-
Comprehensive molecular profiling of combined hepatocellular carcinoma and cholangiocarcinoma reveals distinct Notch signaling subgroups with prognostic significance.Virchows Arch. 2025 Jul 10. doi: 10.1007/s00428-025-04172-9. Online ahead of print. Virchows Arch. 2025. PMID: 40637792
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

