Unraveling the influences of sequence and position on yeast uORF activity using massively parallel reporter systems and machine learning
- PMID: 37227054
- PMCID: PMC10259493
- DOI: 10.7554/eLife.69611
Unraveling the influences of sequence and position on yeast uORF activity using massively parallel reporter systems and machine learning
Abstract
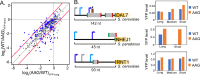
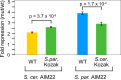
Upstream open-reading frames (uORFs) are potent cis-acting regulators of mRNA translation and nonsense-mediated decay (NMD). While both AUG- and non-AUG initiated uORFs are ubiquitous in ribosome profiling studies, few uORFs have been experimentally tested. Consequently, the relative influences of sequence, structural, and positional features on uORF activity have not been determined. We quantified thousands of yeast uORFs using massively parallel reporter assays in wildtype and ∆upf1 yeast. While nearly all AUG uORFs were robust repressors, most non-AUG uORFs had relatively weak impacts on expression. Machine learning regression modeling revealed that both uORF sequences and locations within transcript leaders predict their effect on gene expression. Indeed, alternative transcription start sites highly influenced uORF activity. These results define the scope of natural uORF activity, identify features associated with translational repression and NMD, and suggest that the locations of uORFs in transcript leaders are nearly as predictive as uORF sequences.
Keywords: S. cerevisiae; genetics; genomics; mRNA Translation; nonsense-mediated decay; uORF.
© 2023, May et al.
Conflict of interest statement
GM, CA, MA, JM, JW, JM No competing interests declared
Figures

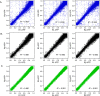



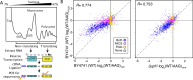
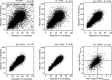
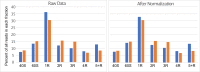





Update of
- doi: 10.1101/2021.04.16.440232
Similar articles
-
High-Throughput Quantitation of Yeast uORF Regulatory Impacts Using FACS-uORF.Methods Mol Biol. 2022;2404:331-351. doi: 10.1007/978-1-0716-1851-6_18. Methods Mol Biol. 2022. PMID: 34694618 Free PMC article.
-
Conserved non-AUG uORFs revealed by a novel regression analysis of ribosome profiling data.Genome Res. 2018 Feb;28(2):214-222. doi: 10.1101/gr.221507.117. Epub 2017 Dec 18. Genome Res. 2018. PMID: 29254944 Free PMC article.
-
Plant transcripts with long or structured upstream open reading frames in the NDL2 5' UTR can escape nonsense-mediated mRNA decay in a reinitiation-independent manner.J Exp Bot. 2023 Jan 1;74(1):91-103. doi: 10.1093/jxb/erac385. J Exp Bot. 2023. PMID: 36169317
-
The regulatory potential of upstream open reading frames in eukaryotic gene expression.Wiley Interdiscip Rev RNA. 2014 Nov-Dec;5(6):765-78. doi: 10.1002/wrna.1245. Epub 2014 Jul 3. Wiley Interdiscip Rev RNA. 2014. PMID: 24995549 Review.
-
A helicase links upstream ORFs and RNA structure.Curr Genet. 2019 Apr;65(2):453-456. doi: 10.1007/s00294-018-0911-z. Epub 2018 Nov 27. Curr Genet. 2019. PMID: 30483885 Free PMC article. Review.
Cited by
-
Deciphering the landscape of cis-acting sequences in natural yeast transcript leaders.Nucleic Acids Res. 2025 Feb 27;53(5):gkaf165. doi: 10.1093/nar/gkaf165. Nucleic Acids Res. 2025. PMID: 40071932 Free PMC article.
-
Predicting the translation efficiency of messenger RNA in mammalian cells.bioRxiv [Preprint]. 2025 Jan 18:2024.08.11.607362. doi: 10.1101/2024.08.11.607362. bioRxiv. 2025. Update in: Nat Biotechnol. 2025 Jul 25. doi: 10.1038/s41587-025-02712-x. PMID: 39149337 Free PMC article. Updated. Preprint.
-
Multilevel gene expression changes in lineages containing adaptive copy number variants.bioRxiv [Preprint]. 2024 Jul 9:2023.10.20.563336. doi: 10.1101/2023.10.20.563336. bioRxiv. 2024. Update in: Mol Biol Evol. 2025 Feb 03;42(2):msaf005. doi: 10.1093/molbev/msaf005. PMID: 37961325 Free PMC article. Updated. Preprint.
-
Tiny but mighty: Diverse functions of uORFs that regulate gene expression.Comput Struct Biotechnol J. 2024 Oct 28;23:3771-3779. doi: 10.1016/j.csbj.2024.10.042. eCollection 2024 Dec. Comput Struct Biotechnol J. 2024. PMID: 39525088 Free PMC article. Review.
-
The regulatory landscape of 5' UTRs in translational control during zebrafish embryogenesis.bioRxiv [Preprint]. 2023 Nov 23:2023.11.23.568470. doi: 10.1101/2023.11.23.568470. bioRxiv. 2023. Update in: Dev Cell. 2025 May 19;60(10):1498-1515.e8. doi: 10.1016/j.devcel.2024.12.038. PMID: 38045294 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

