Microenvironments Matter: Advances in Brain-on-Chip
- PMID: 37232912
- PMCID: PMC10216565
- DOI: 10.3390/bios13050551
Microenvironments Matter: Advances in Brain-on-Chip
Abstract
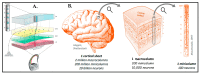
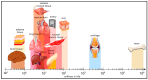
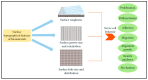
To highlight the particular needs with respect to modeling the unique and complex organization of the human brain structure, we reviewed the state-of-the-art in devising brain models with engineered instructive microenvironments. To acquire a better perspective on the brain's working mechanisms, we first summarize the importance of regional stiffness gradients in brain tissue, varying per layer and the cellular diversities of the layers. Through this, one can acquire an understanding of the essential parameters in emulating the brain in vitro. In addition to the brain's organizational architecture, we addressed also how the mechanical properties have an impact on neuronal cell responses. In this respect, advanced in vitro platforms emerged and profoundly changed the methods of brain modeling efforts from the past, mainly focusing on animal or cell line research. The main challenges in imitating features of the brain in a dish are with regard to composition and functionality. In neurobiological research, there are now methods that aim to cope with such challenges by the self-assembly of human-derived pluripotent stem cells (hPSCs), i.e., brainoids. Alternatively, these brainoids can be used stand-alone or in conjunction with Brain-on-Chip (BoC) platform technology, 3D-printed gels, and other types of engineered guidance features. Currently, advanced in vitro methods have made a giant leap forward regarding cost-effectiveness, ease-of-use, and availability. We bring these recent developments together into one review. We believe our conclusions will give a novel perspective towards advancing instructive microenvironments for BoCs and the understanding of the brain's cellular functions either in modeling healthy or diseased states of the brain.
Keywords: Brain-on-Chip; instructive microenvironment; microfabrication.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



















Similar articles
-
Biosensor integrated brain-on-a-chip platforms: Progress and prospects in clinical translation.Biosens Bioelectron. 2023 Apr 1;225:115100. doi: 10.1016/j.bios.2023.115100. Epub 2023 Jan 24. Biosens Bioelectron. 2023. PMID: 36709589 Review.
-
Next generation human skin constructs as advanced tools for drug development.Exp Biol Med (Maywood). 2017 Nov;242(17):1657-1668. doi: 10.1177/1535370217712690. Epub 2017 Jun 7. Exp Biol Med (Maywood). 2017. PMID: 28592171 Free PMC article. Review.
-
Brain-on-a-Chip: Characterizing the next generation of advanced in vitro platforms for modeling the central nervous system.APL Bioeng. 2021 Jul 30;5(3):030902. doi: 10.1063/5.0055812. eCollection 2021 Sep. APL Bioeng. 2021. PMID: 34368601 Free PMC article.
-
Microfluidic organ-on-chip technology for blood-brain barrier research.Tissue Barriers. 2016 Jan 28;4(1):e1142493. doi: 10.1080/21688370.2016.1142493. eCollection 2016 Jan-Mar. Tissue Barriers. 2016. PMID: 27141422 Free PMC article. Review.
-
Fitting tissue chips and microphysiological systems into the grand scheme of medicine, biology, pharmacology, and toxicology.Exp Biol Med (Maywood). 2017 Oct;242(16):1559-1572. doi: 10.1177/1535370217732765. Exp Biol Med (Maywood). 2017. PMID: 29065799 Free PMC article.
Cited by
-
From Molecules to Mind: The Critical Role of Chitosan, Collagen, Alginate, and Other Biopolymers in Neuroprotection and Neurodegeneration.Molecules. 2025 Feb 22;30(5):1017. doi: 10.3390/molecules30051017. Molecules. 2025. PMID: 40076240 Free PMC article. Review.
-
Navigating the Intersection of Technology and Depression Precision Medicine.Adv Exp Med Biol. 2024;1456:401-426. doi: 10.1007/978-981-97-4402-2_20. Adv Exp Med Biol. 2024. PMID: 39261440 Review.
-
Tailoring epilepsy treatment: personalized micro-physiological systems illuminate individual drug responses.Ann Med Surg (Lond). 2024 Apr 16;86(6):3557-3567. doi: 10.1097/MS9.0000000000002078. eCollection 2024 Jun. Ann Med Surg (Lond). 2024. PMID: 38846814 Free PMC article. Review.
-
The rise of 3D bioprinting advancements in modeling neurodegenerative diseases.Ibrain. 2025 Apr 22;11(2):259-267. doi: 10.1002/ibra.12196. eCollection 2025 Summer. Ibrain. 2025. PMID: 40546880 Free PMC article.
-
Mechanodynamic brain on chip for studying human stem cell derived neuronal networks.Sci Rep. 2025 Aug 13;15(1):29631. doi: 10.1038/s41598-025-14187-6. Sci Rep. 2025. PMID: 40804068 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

