Silver Nanoparticle Targets Fabricated Using Chemical Vapor Deposition Method for Differentiation of Bacteria Based on Lipidomic Profiles in Laser Desorption/Ionization Mass Spectrometry
- PMID: 37237776
- PMCID: PMC10215128
- DOI: 10.3390/antibiotics12050874
Silver Nanoparticle Targets Fabricated Using Chemical Vapor Deposition Method for Differentiation of Bacteria Based on Lipidomic Profiles in Laser Desorption/Ionization Mass Spectrometry
Abstract
The global threat of numerous infectious diseases creates a great need to develop new diagnostic methods to facilitate the appropriate prescription of antimicrobial therapy. More recently, the possibility of using bacterial lipidome analysis via laser desorption/ionization mass spectrometry (LDI-MS) as useful diagnostic tool for microbial identification and rapid drug susceptibility has received particular attention because lipids are present in large quantities and can be easily extracted similar to ribosomal proteins. Therefore, the main goal of the study was to evaluate the efficacy of two different LDI techniques-matrix-assisted (MALDI) and surface-assisted (SALDI) approaches-in the classification of the closely related Escherichia coli strains under cefotaxime addition. Bacterial lipids profiles obtained by using the MALDI technique with different matrices as well as silver nanoparticle (AgNP) targets fabricated using the chemical vapor deposition method (CVD) of different AgNP sizes were analyzed by the means of different multivariate statistical methods such as principal component analysis (PCA), partial least squares discriminant analysis (PLS-DA), sparse partial least squares discriminant analysis (sPLS-DA), and orthogonal projections to latent structures discriminant analysis (OPLS-DA). The analysis showed that the MALDI classification of strains was hampered by interference from matrix-derived ions. In contrast, the lipid profiles generated by the SALDI technique had lower background noise and more signals associated with the sample, allowing E. coli to be successfully classified into cefotaxime-resistant and cefotaxime-sensitive strains, regardless of the size of the AgNPs. AgNP substrates obtained using the CVD method were used for the first time for distinguishing closely related bacterial strains based on their lipidomic profiles and demonstrate high potential as a future diagnostic tool for the detection of antibiotic susceptibility.
Keywords: bacteria; chemical vapor deposition; lipids; mass spectrometry; silver nanoparticles.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
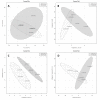
Figures





Similar articles
-
Advances in LDI-MS Analysis: The Role of Chemical Vapor Deposition-Synthesized Silver Nanoparticles in Enhancing Detection of Low-Molecular-Weight Biomolecules.J Am Soc Mass Spectrom. 2024 Sep 4;35(9):2041-2055. doi: 10.1021/jasms.4c00071. Epub 2024 Aug 14. J Am Soc Mass Spectrom. 2024. PMID: 39140654 Free PMC article.
-
Differentiation of Escherichia coli and Shigella flexneri by Metabolite Profiles Obtained Using Gold Nanoparticles-Based Surface-Assisted Laser Desorption/Ionization Mass Spectrometry.Pathogens. 2024 Dec 30;14(1):19. doi: 10.3390/pathogens14010019. Pathogens. 2024. PMID: 39860979 Free PMC article.
-
Bacterial identification by lipid profiling using liquid atmospheric pressure matrix-assisted laser desorption/ionization mass spectrometry.Clin Chem Lab Med. 2020 Jun 25;58(6):930-938. doi: 10.1515/cclm-2019-0908. Clin Chem Lab Med. 2020. PMID: 31851611
-
Nanoparticle-based laser desorption/ionization mass spectrometric analysis of drugs and metabolites.J Food Drug Anal. 2018 Oct;26(4):1215-1228. doi: 10.1016/j.jfda.2018.07.001. Epub 2018 Aug 14. J Food Drug Anal. 2018. PMID: 30249320 Free PMC article. Review.
-
Surface-assisted laser desorption ionization mass spectrometry techniques for application in forensics.Mass Spectrom Rev. 2015 Nov-Dec;34(6):627-40. doi: 10.1002/mas.21431. Epub 2014 Jun 11. Mass Spectrom Rev. 2015. PMID: 24916100 Review.
Cited by
-
Silver Nanoparticles: Synthesis, Structure, Properties and Applications.Nanomaterials (Basel). 2024 Aug 31;14(17):1425. doi: 10.3390/nano14171425. Nanomaterials (Basel). 2024. PMID: 39269087 Free PMC article. Review.
-
Synthesis of Silver Nanoparticles by Chemical Vapor Deposition Method and Its Application in Laser Desorption/Ionization Techniques.Nanomaterials (Basel). 2025 Jun 23;15(13):973. doi: 10.3390/nano15130973. Nanomaterials (Basel). 2025. PMID: 40648680 Free PMC article. Review.
-
Advances in genotypic antimicrobialresistance testing: a comprehensive review.Sci China Life Sci. 2025 Jan;68(1):130-143. doi: 10.1007/s11427-023-2570-4. Epub 2024 Sep 18. Sci China Life Sci. 2025. PMID: 39300049 Review.
-
MALDI-TOF as a powerful tool for identifying and differentiating closely related microorganisms: the strange case of three reference strains of Paenibacillus polymyxa.Sci Rep. 2024 Jan 31;14(1):2585. doi: 10.1038/s41598-023-50010-w. Sci Rep. 2024. PMID: 38297004 Free PMC article.
-
Nanosilver: An Old Antibacterial Agent with Great Promise in the Fight against Antibiotic Resistance.Antibiotics (Basel). 2023 Jul 31;12(8):1264. doi: 10.3390/antibiotics12081264. Antibiotics (Basel). 2023. PMID: 37627684 Free PMC article. Review.
References
-
- Liang T., Leung L.M., Opene B., Fondrie W.E., Lee Y.I., Chandler C.E., Yoon S.H., Doi Y., Ernst R.K., Goodlett D.R. Rapid Microbial Identification and Antibiotic Resistance Detection by Mass Spectrometric Analysis of Membrane Lipids. Anal. Chem. 2019;91:1286–1294. doi: 10.1021/acs.analchem.8b02611. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

