An In Vitro Model of Glioma Development
- PMID: 37239349
- PMCID: PMC10217752
- DOI: 10.3390/genes14050990
An In Vitro Model of Glioma Development
Abstract
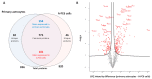
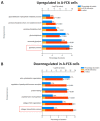
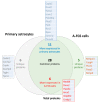
Gliomas are the prevalent forms of brain cancer and derive from glial cells. Among them, astrocytomas are the most frequent. Astrocytes are fundamental for most brain functions, as they contribute to neuronal metabolism and neurotransmission. When they acquire cancer properties, their functions are altered, and, in addition, they start invading the brain parenchyma. Thus, a better knowledge of transformed astrocyte molecular properties is essential. With this aim, we previously developed rat astrocyte clones with increasing cancer properties. In this study, we used proteomic analysis to compare the most transformed clone (A-FC6) with normal primary astrocytes. We found that 154 proteins are downregulated and 101 upregulated in the clone. Moreover, 46 proteins are only expressed in the clone and 82 only in the normal cells. Notably, only 11 upregulated/unique proteins are encoded in the duplicated q arm of isochromosome 8 (i(8q)), which cytogenetically characterizes the clone. Since both normal and transformed brain cells release extracellular vesicles (EVs), which might induce epigenetic modifications in the neighboring cells, we also compared EVs released from transformed and normal astrocytes. Interestingly, we found that the clone releases EVs containing proteins, such as matrix metalloproteinase 3 (MMP3), that can modify the extracellular matrix, thus allowing invasion.
Keywords: astrocyte cell lines; astrocytomas; chromosome alterations; epigenetic alterations; extracellular vesicles (EVs); metalloproteinases; proteomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

