Protein Fitness Prediction Is Impacted by the Interplay of Language Models, Ensemble Learning, and Sampling Methods
- PMID: 37242577
- PMCID: PMC10224321
- DOI: 10.3390/pharmaceutics15051337
Protein Fitness Prediction Is Impacted by the Interplay of Language Models, Ensemble Learning, and Sampling Methods
Abstract
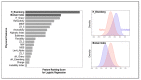
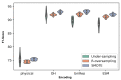
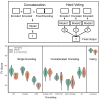
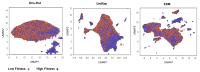
Advances in machine learning (ML) and the availability of protein sequences via high-throughput sequencing techniques have transformed the ability to design novel diagnostic and therapeutic proteins. ML allows protein engineers to capture complex trends hidden within protein sequences that would otherwise be difficult to identify in the context of the immense and rugged protein fitness landscape. Despite this potential, there persists a need for guidance during the training and evaluation of ML methods over sequencing data. Two key challenges for training discriminative models and evaluating their performance include handling severely imbalanced datasets (e.g., few high-fitness proteins among an abundance of non-functional proteins) and selecting appropriate protein sequence representations (numerical encodings). Here, we present a framework for applying ML over assay-labeled datasets to elucidate the capacity of sampling techniques and protein encoding methods to improve binding affinity and thermal stability prediction tasks. For protein sequence representations, we incorporate two widely used methods (One-Hot encoding and physiochemical encoding) and two language-based methods (next-token prediction, UniRep; masked-token prediction, ESM). Elaboration on performance is provided over protein fitness, protein size, and sampling techniques. In addition, an ensemble of protein representation methods is generated to discover the contribution of distinct representations and improve the final prediction score. We then implement multiple criteria decision analysis (MCDA; TOPSIS with entropy weighting), using multiple metrics well-suited for imbalanced data, to ensure statistical rigor in ranking our methods. Within the context of these datasets, the synthetic minority oversampling technique (SMOTE) outperformed undersampling while encoding sequences with One-Hot, UniRep, and ESM representations. Moreover, ensemble learning increased the predictive performance of the affinity-based dataset by 4% compared to the best single-encoding candidate (F1-score = 97%), while ESM alone was rigorous enough in stability prediction (F1-score = 92%).
Keywords: MCDA; TOPSIS; embeddings; ensemble learning; imbalanced assay-labeled datasets; machine learning; protein fitness prediction; sampling methods; sequence representation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



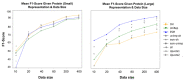
References
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

