De novo transcriptome assembly and annotation for gene discovery in Salamandra salamandra at the larval stage
- PMID: 37244908
- PMCID: PMC10224929
- DOI: 10.1038/s41597-023-02217-9
De novo transcriptome assembly and annotation for gene discovery in Salamandra salamandra at the larval stage
Abstract
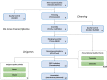
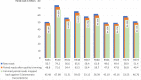
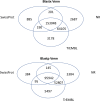
Dispersal is a key process in ecology and evolutionary biology, as it shapes biodiversity patterns over space and time. Attitude to disperse is unevenly distributed among individuals within populations, and that individual personality can have pivotal roles in the shaping of this attitude. Here, we assembled and annotated the first de novo transcriptome of the head tissues of Salamandra salamandra from individuals, representative of distinct behavioral profiles. We obtained 1,153,432,918 reads, which were successfully assembled and annotated. The high-quality of the assembly was confirmed by three assembly validators. The alignment of contigs against the de novo transcriptome led to a mapping percentage higher than 94%. The homology annotation with DIAMOND led to 153,048 (blastx) and 95,942 (blastp) shared contigs, annotated on NR, Swiss-Prot and TrEMBL. The domain and site protein prediction led to 9850 GO-annotated contigs. This de novo transcriptome represents reliable reference for comparative gene expression studies between alternative behavioral types, for comparative gene expression studies within Salamandra, and for whole transcriptome and proteome studies in amphibians.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Brain de novo transcriptome assembly of a toad species showing polymorphic anti-predatory behavior.Sci Data. 2022 Oct 13;9(1):619. doi: 10.1038/s41597-022-01724-5. Sci Data. 2022. PMID: 36229462 Free PMC article.
-
De Novo Assembly and Characterization of the Transcriptome of Grasshopper Shirakiacris shirakii.Int J Mol Sci. 2016 Jul 22;17(7):1110. doi: 10.3390/ijms17071110. Int J Mol Sci. 2016. PMID: 27455245 Free PMC article.
-
A high-quality annotated transcriptome of swine peripheral blood.BMC Genomics. 2017 Jun 24;18(1):479. doi: 10.1186/s12864-017-3863-7. BMC Genomics. 2017. PMID: 28646867 Free PMC article.
-
Transcriptome Analysis of the Emerald Ash Borer (EAB), Agrilus planipennis: De Novo Assembly, Functional Annotation and Comparative Analysis.PLoS One. 2015 Aug 5;10(8):e0134824. doi: 10.1371/journal.pone.0134824. eCollection 2015. PLoS One. 2015. PMID: 26244979 Free PMC article.
-
A simple guide to de novo transcriptome assembly and annotation.Brief Bioinform. 2022 Mar 10;23(2):bbab563. doi: 10.1093/bib/bbab563. Brief Bioinform. 2022. PMID: 35076693 Free PMC article. Review.
Cited by
-
Use of Field pathogenomics approach for Puccinia striiformis f. sp. tritici race identification and phylogenomic delineation in North India.World J Microbiol Biotechnol. 2025 May 6;41(5):166. doi: 10.1007/s11274-025-04391-x. World J Microbiol Biotechnol. 2025. PMID: 40325275
-
Integrated de novo transcriptome of Culex pipiens mosquito larvae as a resource for genetic control strategies.Sci Data. 2024 May 9;11(1):471. doi: 10.1038/s41597-024-03285-1. Sci Data. 2024. PMID: 38724521 Free PMC article.
-
De novo transcriptome assembly of an Antarctic nematode for the study of thermal adaptation in marine parasites.Sci Data. 2023 Oct 19;10(1):720. doi: 10.1038/s41597-023-02591-4. Sci Data. 2023. PMID: 37857654 Free PMC article.
-
HPC-T-Assembly: a pipeline for de novo transcriptome assembly of large multi-specie datasets.BMC Bioinformatics. 2025 Apr 28;26(1):113. doi: 10.1186/s12859-025-06121-4. BMC Bioinformatics. 2025. PMID: 40295976 Free PMC article.
-
De novo transcriptome assembly of the Mediterranean sea-rock pool mosquitoes Aedes mariae and Aedes zammitii.Sci Data. 2025 Jan 20;12(1):115. doi: 10.1038/s41597-025-04393-2. Sci Data. 2025. PMID: 39833234 Free PMC article.
References
-
- Clobert, J., Baguette, M., Benton, T. G. & Bullock, J. M. Dispersal ecology and evolution. (Oxford University Press. 462 pp. - 2012)
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

