Distribution and diversity of 'Tectomicrobia', a deep-branching uncultivated bacterial lineage harboring rich producers of bioactive metabolites
- PMID: 37248312
- PMCID: PMC10227082
- DOI: 10.1038/s43705-023-00259-z
Distribution and diversity of 'Tectomicrobia', a deep-branching uncultivated bacterial lineage harboring rich producers of bioactive metabolites
Abstract
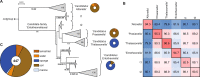
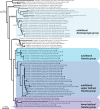
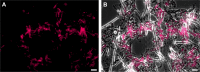
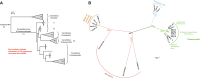
Genomic and functional analyses of bacterial sponge symbionts belonging to the uncultivated candidate genus 'Entotheonella' has revealed them as the prolific producers of bioactive compounds previously identified from their invertebrate hosts. These studies also suggested 'Entotheonella' as the first members of a new candidate phylum, 'Tectomicrobia'. Here we analyzed the phylogenetic structure and environmental distribution of this as-yet sparsely populated phylum-like lineage. The data show that 'Entotheonella' and other 'Tectomicrobia' are not restricted to marine habitats but widely distributed among terrestrial locations. The inferred phylogenetic trees suggest several intra-phylum lineages with diverse lifestyles. Of these, the previously described 'Entotheonella' lineage can be more accurately divided into at least three different candidate genera with the terrestrial 'Candidatus Prasianella', the largely terrestrial 'Candidatus Allonella', the 'Candidatus Thalassonella' comprising sponge-associated members, and the more widely distributed 'Candidatus Entotheonella'. Genomic characterization of 'Thalassonella' members from a range of sponge hosts did not suggest a role as providers of natural products, despite high genomic similarity to 'Entotheonella' regarding primary metabolism and implied lifestyle. In contrast, the analysis revealed a correlation between the revised 'Entotheonella' 16S rRNA gene phylogeny and a specific association with sponges and their natural products. This feature might serve as a discovery method to accelerate the identification of new chemically rich 'Entotheonella' variants, and led to the identification of the first 'Entotheonella' symbiont in a non-tetractinellid sponge, Psammocinia sp., indicating a wide host distribution of 'Entotheonella'-based chemical symbiosis.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


References
Grants and funding
- R01CA47135/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- R01 CA047135/CA/NCI NIH HHS/United States
- ETH-21 18-2/Eidgenössische Technische Hochschule Zürich (Federal Institute of Technology Zurich)
- 205320_185077/Schweizerischer Nationalfonds zur Förderung der Wissenschaftlichen Forschung (Swiss National Science Foundation)
- doi.org/10.37807/GBMF9204/Gordon and Betty Moore Foundation (Gordon E. and Betty I. Moore Foundation)
- 679849/EC | Horizon 2020 Framework Programme (EU Framework Programme for Research and Innovation H2020)
- 897121/EC | EU Framework Programme for Research and Innovation H2020 | H2020 Priority Excellent Science | H2020 Marie Skłodowska-Curie Actions (H2020 Excellent Science - Marie Skłodowska-Curie Actions)

