Comprehensive Membrane N-Glycoproteomics Using Human Breast Cancer Cell Line Pairs
- PMID: 37250596
- PMCID: PMC10213999
- DOI: 10.5702/massspectrometry.A0117
Comprehensive Membrane N-Glycoproteomics Using Human Breast Cancer Cell Line Pairs
Abstract
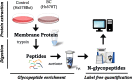
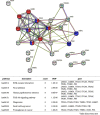
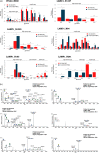
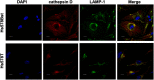
Aberrant glycosylation of membrane proteins is a hallmark of cancer and a useful molecular marker for the diagnosis of breast cancer (BC). However, the molecular mechanisms by which altered glycosylation affects the malignant transformations associated with BC are poorly understood. Accordingly, we performed comparative membrane N-glycoproteomics using the human BC cell line pair, Hs578T, and its syngeneic normal cell line, Hs578Bst. A total of 359 N-glycoforms derived from 113 proteins were identified in both cell lines, of which 27 were found only in Hs578T cells. Significant changes in N-glycosylation were found in the lysosome-associated membrane protein 1 (LAMP1), the integrin family, and laminin. Confocal immunofluorescence microscopy images revealed the accumulation of lysosomes in the perinuclear space in cancer cells, which could be associated with marked changes in LAMP1 glycosylation, such as a decreased level of polylactosamine chains. Overall, the alterations in glycosylation may be involved in changes in the adhesion and degradation of BC cells.
Keywords: LC/MS/MS; N-glycoproteomics; glycosylation; membrane glycoproteins.
Copyright © 2023 Daisuke Takakura, Haruka Yoshida, Shoko Ohashi, and Nana Kawasaki.
Figures


References
-
- Y. Z. Jiang, D. Ma, C. Suo, J. Shi, M. Xue, X. Hu, Y. Xiao, K. D. Yu, Y. R. Liu, Y. Yu, Y. Zheng, X. Li, C. Zhang, P. Hu, J. Zhang, Q. Hua, J. Zhang, W. Hou, L. Ren, D. Bao, B. Li, J. Yang, L. Yao, W. J. Zuo, S. Zhao, Y. Gong, Y. X. Ren, Y. X. Zhao, Y. S. Yang, Z. Niu, Z. G. Cao, D. G. Stover, C. Verschraegen, V. Kaklamani, A. Daemen, J. R. Benson, K. Takabe, F. Bai, D. Q. Li, P. Wang, L. Shi, W. Huang, Z. M. Shao. Genomic and transcriptomic landscape of triple-negative breast cancers: Subtypes and treatment strategies. Cancer Cell 35: 428–440.e5, 2019. - PubMed
-
- P. Jézéquel, O. Kerdraon, H. Hondermarck, C. Guérin-Charbonnel, H. Lasla, W. Gouraud, J. L. Canon, A. Gombos, F. Dalenc, S. Delaloge, J. Lemonnier, D. Loussouarn, V. Verrièle, M. Campone. Identification of three subtypes of triple-negative breast cancer with potential therapeutic implications. Breast Cancer Res. 21: 65, 2019. - PMC - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

