Detection and Separation of DNA and Silver Nanoparticles Using a Solid-State Nanopore
- PMID: 37251189
- PMCID: PMC10210173
- DOI: 10.1021/acsomega.3c00152
Detection and Separation of DNA and Silver Nanoparticles Using a Solid-State Nanopore
Abstract
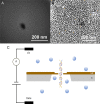
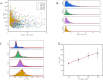
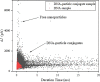
Nanopore sensors, a new generation of single-molecule sensors, are increasingly used to detect and analyze various analytes and have great potential for rapid gene sequencing. However, there are still some problems in the preparation of small diameter nanopores, such as imprecise pore size and porous defects, while the detection accuracy of large-diameter nanopores is relatively low. Therefore, how to achieve more precise detection of large diameter nanopore sensors is an urgent problem to be studied. Here, SiN nanopore sensors were used to detect DNA molecules and silver nanoparticles (NPs) separately and in combination. The experimental results show that large-size solid-state nanopore sensors can identify and discriminate between DNA molecules, NPs, and NP-bound DNA molecules clearly according to resistive pulses. In addition, the detection mechanism of using NPs to assist in identifying target DNA molecules in this study is different from previous reports. We find that silver NPs can simultaneously bind to multiple probes and target DNA molecules and generate a larger blocking current than free DNA molecules when passing through the nanopore. In conclusion, our research indicates that large-sized nanopores can distinguish the translocation events, thereby identifying the presence of the target DNA molecules in the sample. This nanopore-sensing platform can produce rapid and accurate nucleic acid detection. Its application in medical diagnosis, gene therapy, virus identification, and many other fields is highly significant.
© 2023 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures



Similar articles
-
Recent advances of small molecule detection in nanopore sensing.Talanta. 2024 Sep 1;277:126323. doi: 10.1016/j.talanta.2024.126323. Epub 2024 May 25. Talanta. 2024. PMID: 38810384 Review.
-
Solid-State Nanopore Single-Molecule Sensing of DNAzyme Cleavage Reaction Assisted with Nucleic Acid Nanostructure.ACS Appl Mater Interfaces. 2018 Aug 8;10(31):26555-26565. doi: 10.1021/acsami.8b09505. Epub 2018 Jul 26. ACS Appl Mater Interfaces. 2018. PMID: 30016075
-
Solid-State Nanopore Sensors with Enhanced Sensitivity through Nucleic Acid Amplification.Anal Chem. 2023 Nov 28;95(47):17153-17161. doi: 10.1021/acs.analchem.3c03806. Epub 2023 Nov 15. Anal Chem. 2023. PMID: 37966312 Review.
-
Nanoparticle-assisted detection of nucleic acids in a polymeric nanopore with a large pore size.Biosens Bioelectron. 2022 Jan 15;196:113697. doi: 10.1016/j.bios.2021.113697. Epub 2021 Oct 8. Biosens Bioelectron. 2022. PMID: 34649096 Free PMC article.
-
Stable fabrication of a large nanopore by controlled dielectric breakdown in a high-pH solution for the detection of various-sized molecules.Sci Rep. 2019 Sep 11;9(1):13143. doi: 10.1038/s41598-019-49622-y. Sci Rep. 2019. PMID: 31511597 Free PMC article.
References
-
- Kumar H.; Lansac Y.; Glaser M. A.; Maiti P. K. Biopolymers in nanopores: challenges and opportunities. Soft Matter 2011, 7, 5898–5907. 10.1039/c0sm01517b. - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

