The CCP4 suite: integrative software for macromolecular crystallography
- PMID: 37259835
- PMCID: PMC10233625
- DOI: 10.1107/S2059798323003595
The CCP4 suite: integrative software for macromolecular crystallography
Abstract
The Collaborative Computational Project No. 4 (CCP4) is a UK-led international collective with a mission to develop, test, distribute and promote software for macromolecular crystallography. The CCP4 suite is a multiplatform collection of programs brought together by familiar execution routines, a set of common libraries and graphical interfaces. The CCP4 suite has experienced several considerable changes since its last reference article, involving new infrastructure, original programs and graphical interfaces. This article, which is intended as a general literature citation for the use of the CCP4 software suite in structure determination, will guide the reader through such transformations, offering a general overview of the new features and outlining future developments. As such, it aims to highlight the individual programs that comprise the suite and to provide the latest references to them for perusal by crystallographers around the world.
Keywords: CCP4; Collaborative Computational Project No. 4; crystallography software; macromolecular crystallography.
open access.
Figures

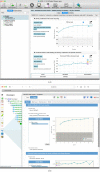
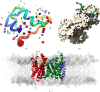
References
-
- Agirre, J., Iglesias-Fernández, J., Rovira, C., Davies, G. J., Wilson, K. S. & Cowtan, K. D. (2015). Nat. Struct. Mol. Biol. 22, 833–834. - PubMed
-
- Allen, F. H. & Bruno, I. J. (2010). Acta Cryst. B66, 380–386. - PubMed
-
- Armstrong, D. R., Berrisford, J. M., Conroy, M. J., Gutmanas, A., Anyango, S., Choudhary, P., Clark, A. R., Dana, J. M., Deshpande, M., Dunlop, R., Gane, P., Gáborová, R., Gupta, D., Haslam, P., Koča, J., Mak, L., Mir, S., Mukhopadhyay, A., Nadzirin, N., Nair, S., Paysan-Lafosse, T., Pravda, L., Sehnal, D., Salih, O., Smart, O., Tolchard, J., Varadi, M., Svobodova-Vařeková, R., Zaki, H., Kleywegt, G. J. & Velankar, S. (2020). Nucleic Acids Res. 48, D335–D343. - PMC - PubMed
MeSH terms
Substances
Grants and funding
- BB/P012523/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/L006383/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/S005099/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/S007083/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- 200898/WT_/Wellcome Trust/United Kingdom
- BB/F020805/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/T012935/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/S007105/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- 203149/WT_/Wellcome Trust/United Kingdom
- BB/P012574/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/S007040/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- 209407/Z/17/Z/WT_/Wellcome Trust/United Kingdom
- MC_UP_A025_1012/MRC_/Medical Research Council/United Kingdom
- WT_/Wellcome Trust/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

