Genome-Wide Association Study of CKD Progression
- PMID: 37261792
- PMCID: PMC10482057
- DOI: 10.1681/ASN.0000000000000170
Genome-Wide Association Study of CKD Progression
Abstract
Significance statement: Rapid progression of CKD is associated with poor clinical outcomes. Most previous studies looking for genetic factors associated with low eGFR have used cross-sectional data. The authors conducted a meta-analysis of genome-wide association studies of eGFR decline among 116,870 participants with CKD, focusing on longitudinal data. They identified three loci (two of them novel) associated with longitudinal eGFR decline. In addition to the known UMOD/PDILT locus, variants within BICC1 were associated with significant differences in longitudinal eGFR slope. Variants within HEATR4 also were associated with differences in eGFR decline, but only among Black/African American individuals without diabetes. These findings help characterize molecular mechanisms of eGFR decline in CKD and may inform new therapeutic approaches for progressive kidney disease.
Background: Rapid progression of CKD is associated with poor clinical outcomes. Despite extensive study of the genetics of cross-sectional eGFR, only a few loci associated with eGFR decline over time have been identified.
Methods: We performed a meta-analysis of genome-wide association studies of eGFR decline among 116,870 participants with CKD-defined by two outpatient eGFR measurements of <60 ml/min per 1.73 m 2 , obtained 90-365 days apart-from the Million Veteran Program and Vanderbilt University Medical Center's DNA biobank. The primary outcome was the annualized relative slope in outpatient eGFR. Analyses were stratified by ethnicity and diabetes status and meta-analyzed thereafter.
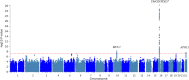
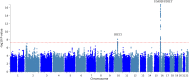
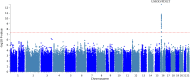
Results: In cross-ancestry meta-analysis, the strongest association was rs77924615, near UMOD / PDILT ; each copy of the G allele was associated with a 0.30%/yr faster eGFR decline ( P = 4.9×10 -27 ). We also observed an association within BICC1 (rs11592748), where every additional minor allele was associated with a 0.13%/yr slower eGFR decline ( P = 5.6×10 -9 ). Among participants without diabetes, the strongest association was the UMOD/PDILT variant rs36060036, associated with a 0.27%/yr faster eGFR decline per copy of the C allele ( P = 1.9×10 -17 ). Among Black participants, a significantly faster eGFR decline was associated with variant rs16996674 near APOL1 (R 2 =0.29 with the G1 high-risk genotype); among Black participants with diabetes, lead variant rs11624911 near HEATR4 also was associated with a significantly faster eGFR decline. We also nominally replicated loci with known associations with eGFR decline, near PRKAG2, FGF5, and C15ORF54.
Conclusions: Three loci were significantly associated with longitudinal eGFR change at genome-wide significance. These findings help characterize molecular mechanisms of eGFR decline and may contribute to the development of new therapeutic approaches for progressive CKD.
Copyright © 2023 by the American Society of Nephrology.
Conflict of interest statement
A. Bick reports Ownership Interest: TenSixteen Bio. C.P. Chung reports Advisory or Leadership Role: Clinical Pharmacology and Therapeutics, Arthritis Care and Research, and Clinical Rheumatology. A.M. Hung reports Consultancy: NHLBI consultant for Gene and life interaction grant; Research Funding: VHA CSR&D Merit “Genetics of Kidney Disease & Hypertension, Risk Prediction and Drug Response,” Vertex Grant to VUMC; and Advisory or Leadership Role: Co-Chair Million Veteran Program Publications & Presentation committee for Veterans Affairs, Co-chair Pharmacogenomics for COVID-19 Million Veteran Program, Journal of renal nutrition, Section Editor Clinical Nephrology, Standing member SRC HSR&D bioinformatics, Ad-hoc SRC NHLBI, Ad-hoc Scientific Review Committee CSR&D, Ad-Scientific Review Committee KNOD. T.A. Ikizler reports Consultancy: Abbott Renal Care, Fresenius–Kabi, La Renon, Nestle; Honoraria: Fresenius–Kabi, Abbott Renal Care, La Renon, Nestle; Patents or Royalties: Vanderbilt University Medical Center; and Advisory or Leadership Role: Kidney International. L. Lipworth reports Research Funding: EpidStrategies. M.E. Matheny reports Consultancy: NIH-VA-DoD Pain Management Grant Consortium (PMC3); and Advisory or Leadership Role: SMRB Study Section, VA HSR&D, Informatics & Methods Section, Steering Committee—Indianapolis VA HSR&D COIN Center, Steering Committee—VA HSR&D VIREC, Steering Committee—Salt Lake City VA HSR&D COIN Center, and VA ORD Million Veterans Program Executive Steering Committee. C. Robinson-Cohen reports Advisory or Leadership Role:
Figures

Comment in
-
Incorporating Linear Mixed Models into GWAS of Kidney Function Decline.J Am Soc Nephrol. 2023 Sep 1;34(9):1473-1475. doi: 10.1681/ASN.0000000000000175. J Am Soc Nephrol. 2023. PMID: 37656511 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
- U01 HG004798/HG/NHGRI NIH HHS/United States
- UL1 RR024975/RR/NCRR NIH HHS/United States
- U19 HL065962/HL/NHLBI NIH HHS/United States
- I01 CX001897/CX/CSRD VA/United States
- P50 GM115305/GM/NIGMS NIH HHS/United States
- R01 HD074711/HD/NICHD NIH HHS/United States
- R01 NS032830/NS/NINDS NIH HHS/United States
- K12 HD043483/HD/NICHD NIH HHS/United States
- R01 DK122075/DK/NIDDK NIH HHS/United States
- U01 HG006378/HG/NHGRI NIH HHS/United States
- S10 RR025141/RR/NCRR NIH HHS/United States
- UL1 TR002243/TR/NCATS NIH HHS/United States
- UL1 TR000445/TR/NCATS NIH HHS/United States
- RC2 GM092618/GM/NIGMS NIH HHS/United States
- K12 AR084232/AR/NIAMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

