Acquisition and carriage of genetically diverse multi-drug resistant gram-negative bacilli in hospitalised newborns in The Gambia
- PMID: 37270610
- PMCID: PMC10239441
- DOI: 10.1038/s43856-023-00309-6
Acquisition and carriage of genetically diverse multi-drug resistant gram-negative bacilli in hospitalised newborns in The Gambia
Abstract
Background: This detailed genomic study characterised multi-drug resistant-Gram negative bacilli (MDR-GNB) carriage in neonates < 2 kg and paired mothers at a low-resource African hospital.
Methods: This cross-sectional cohort study was conducted at the neonatal referral unit in The Gambia with weekly neonatal skin and peri-anal sampling and paired maternal recto-vaginal swabs. Prospective bacteriological culture used MacConkey agar with species identification by API20E and API20NE. All GNB isolates underwent whole genome sequencing on Illumina Miseq platform. Multi-Locus Sequence Typing and SNP-distance analysis identified strain type and relatedness.
Results: 135 swabs from 34 neonates and 21 paired mothers, yielded 137 GNB isolates, of which 112 are high quality de novo assemblies. Neonatal MDR-GNB carriage prevalence is 41% (14/34) at admission with 85% (11/13) new acquisition by 7d. Multiple MDR and ESBL-GNB species are carried at different timepoints, most frequently K. pneumoniae and E. coli, with heterogeneous strain diversity and no evidence of clonality. 111 distinct antibiotic resistance genes are mostly beta lactamases (Bla-AMPH, Bla-PBP, CTX-M-15, Bla-TEM-105). 76% (16/21) and 62% (13/21) of mothers have recto-vaginal carriage of ≥1 MDR-GNB and ESBL-GNB respectively, mostly MDR-E. coli (76%, 16/21) and MDR-K. pneumoniae (24%, 5/21). Of 21 newborn-mother dyads, only one have genetically identical isolates (E. coli ST131 and K. pneumoniae ST3476).
Conclusions: Gambian hospitalised neonates exhibit high MDR and ESBL-GNB carriage prevalence with acquisition between birth and 7d with limited evidence supporting mother to neonate transmission. Genomic studies in similar settings are required to further understand transmission and inform targeted surveillance and infection prevention policies.
Plain language summary
Bacteria that are resistant to multiple antibiotics are an important cause of infection and death of newborns in low-resource countries, especially small or premature babies born in hospital settings. How these resistant bacteria are acquired on the skin and in the gut of newborns is not known, particularly whether they are commonly transferred from mothers. We studied the bacteria present in small Gambian newborns and their mothers to understand the type of bacteria, amount of antibiotic resistance, number of newborns and mothers affected and similarity of these bacteria between newborns and their mothers. We found that despite many newborns carrying these bacteria, they are different from those present in mothers. This suggests that the bacteria are acquired from the hospital environment. Our study highlights the importance of developing strategies to identify and reduce the presence of such bacteria in hospitals to reduce their acquisition by vulnerable hospitalised newborns.
© 2023. The Author(s).
Conflict of interest statement
The authors declare the following competing interests: B.K. reports grants from the MRC UK Research & Innovation (UKRI), United Kingdom; Wellcome Trust, United Kingdom and Bill and Melinda Gates Foundation (BMGF), United States for a variety of projects related to vaccines and maternal/newborn health. B.K. attended the Gates Global Challenge Meeting in 2022, supported by BMGF, and is also on the Data Safety Monitoring Board for a COVID producing vaccine company. Td.S. is an Editorial Board Member for
Figures

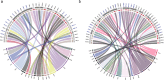

References
-
- United Nations Inter-agency Group for Child Mortality Estimation. Levels & Trends in Child Mortality: Report 2020, Estimates developed by the United Nations Inter-agency Group for Child Mortality Estimation. New York: United Nations Children’s Fund; 2020.
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

