Single-cell transcriptomics suggest distinct upstream drivers of IL-17A/F in hidradenitis versus psoriasis
- PMID: 37271319
- PMCID: PMC11057969
- DOI: 10.1016/j.jaci.2023.05.012
Single-cell transcriptomics suggest distinct upstream drivers of IL-17A/F in hidradenitis versus psoriasis
Abstract
Background: On the basis of the mounting evidence that type 17 T (T17) cells and increased IL-17 play a key role in driving hidradenitis suppurativa (HS) lesion development, biologic agents used previously in psoriasis that block signaling of IL-17A and/or IL-17F isoforms have been repurposed to treat HS.
Objective: Our research aimed to characterize the transcriptome of HS T17 cells compared to the transcriptome of psoriasis T17 cells, along with their ligand-receptor interactions with neighborhood immune cell subsets.
Methods: Single-cell data of 12,300 cutaneous immune cells from 8 deroofing surgical HS skin samples including dermal tunnels were compared to single-cell data of psoriasis skin (19,525 cells from 11 samples) and control skin (11,920 cells from 10 samples). All single-cell data were generated by the same protocol.
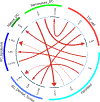
Results: HS T17 cells expressed lower levels of IL23R and higher levels of IL1R1 and IL17F compared to psoriasis T17 cells (P < .05). HS Treg cells expressed higher levels of IL1R1 and IL17F compared to psoriasis Treg cells (P < .05). Semimature dendritic cells were the major immune cell subsets expressing IL1B in HS, and IL-1β ligand-receptor interactions between semimature dendritic cells and T17 cells were increased in HS compared to psoriasis (P < .05). HS dermal tunnel keratinocytes expressed inflammatory cytokines (IL17C, IL1A, IL1B, and IL6) that differed from the HS epidermis keratinocytes (IL36G) (P < .05). IL6, which synergizes with IL1B to maintain cytokine expression in T17 cells, was mainly expressed by fibroblasts in HS, which also expressed IL11+ inflammatory fibroblast genes (IL11, IL24, IL6, and POSTN) involved in the paracrine IL-1/IL-6 loop.
Conclusion: The IL-1β-T17 cell cytokine axis is likely a dominant pathway in HS with HS T17 cells activated by IL-1β signaling, unlike psoriasis T17 cells, which are activated by IL-23 signaling.
Keywords: Hidradenitis suppurativa; IL-1; IL-17A; IL-17F; IL-1R1; IL-23; T cell; dendritic cell; fibroblast; keratinocyte; psoriasis; single-cell RNA sequencing; type 17 T cells.
Published by Elsevier Inc.
Conflict of interest statement
J.K. was supported by NIAMS K23 Career Development Award (K23AR080043). J.K. has received research support from Novartis and AbbVie. MAL has served on the advisory boards for Abbvie, InflaRx, Janssen, and UCB, Viela Bio, and consulted for Almirall, BSN medical, Incyte, Janssen, Kymera, Phoenicis, and XBiotech. MAL is on the medical board of the Hidradenitis Suppurativa Foundation. J.G.K. has received research support from Pfizer, Amgen, Janssen, Lilly, Merck, Novartis, Kadmon, Dermira, Boehringer, Innovaderm, Kyowa, BMS, Serono, BiogenIdec, Delenex, AbbVie, Sanofi, Baxter, Paraxel, Xenoport, and Kineta. The rest of the authors declare that they have no relevant conflict of interest.
Figures





References
-
- Kimball AB, Okun MM, Williams DA, Gottlieb AB, Papp KA, Zouboulis CC, et al. Two phase 3 trials of adalimumab for hidradenitis suppurativa. N Engl J Med. 2016;375:422–34. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

