Genetic architecture of heart mitochondrial proteome influencing cardiac hypertrophy
- PMID: 37276142
- PMCID: PMC10241513
- DOI: 10.7554/eLife.82619
Genetic architecture of heart mitochondrial proteome influencing cardiac hypertrophy
Abstract
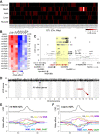
Mitochondria play an important role in both normal heart function and disease etiology. We report analysis of common genetic variations contributing to mitochondrial and heart functions using an integrative proteomics approach in a panel of inbred mouse strains called the Hybrid Mouse Diversity Panel (HMDP). We performed a whole heart proteome study in the HMDP (72 strains, n=2-3 mice) and retrieved 848 mitochondrial proteins (quantified in ≥50 strains). High-resolution association mapping on their relative abundance levels revealed three trans-acting genetic loci on chromosomes (chr) 7, 13 and 17 that regulate distinct classes of mitochondrial proteins as well as cardiac hypertrophy. DAVID enrichment analyses of genes regulated by each of the loci revealed that the chr13 locus was highly enriched for complex-I proteins (24 proteins, P=2.2E-61), the chr17 locus for mitochondrial ribonucleoprotein complex (17 proteins, P=3.1E-25) and the chr7 locus for ubiquinone biosynthesis (3 proteins, P=6.9E-05). Follow-up high resolution regional mapping identified NDUFS4, LRPPRC and COQ7 as the candidate genes for chr13, chr17 and chr7 loci, respectively, and both experimental and statistical analyses supported their causal roles. Furthermore, a large cohort of Diversity Outbred mice was used to corroborate Lrpprc gene as a driver of mitochondrial DNA (mtDNA)-encoded gene regulation, and to show that the chr17 locus is specific to heart. Variations in all three loci were associated with heart mass in at least one of two independent heart stress models, namely, isoproterenol-induced heart failure and diet-induced obesity. These findings suggest that common variations in certain mitochondrial proteins can act in trans to influence tissue-specific mitochondrial functions and contribute to heart hypertrophy, elucidating mechanisms that may underlie genetic susceptibility to heart failure in human populations.
Keywords: computational biology; genetic, association studies; genetics; genomics; heart failure; hypertrophy; metabolic syndrome; mitochondria; mouse; proteomics; systems biology.
© 2023, Chella Krishnan et al.
Conflict of interest statement
KC, EE, MK, SP, LC, AV, IG, CL, YC, CP, KK, VS, DA, MP, CL, MS, NR, GC, AA, BP, AL No competing interests declared, DJ Senior Editor, eLife; Senior Editor for the Systems Genetics Special Issue (DEJ had no involvement in the decision making for this submission)
Figures




Update of
References
-
- Adjobo-Hermans MJW, de Haas R, Willems P, Wojtala A, van Emst-de Vries SE, Wagenaars JA, van den Brand M, Rodenburg RJ, Smeitink JAM, Nijtmans LG, Sazanov LA, Wieckowski MR, Koopman WJH. Ndufs4 deletion triggers loss of Ndufa12 in Ndufs4(-/-) mice and Leigh syndrome patients: A stabilizing role for Ndufaf2. Biochimica et Biophysica Acta. Bioenergetics. 2020;1861:148213. doi: 10.1016/j.bbabio.2020.148213. - DOI - PubMed
-
- Bennett BJ, Farber CR, Orozco L, Kang HM, Ghazalpour A, Siemers N, Neubauer M, Neuhaus I, Yordanova R, Guan B, Truong A, Yang W, He A, Kayne P, Gargalovic P, Kirchgessner T, Pan C, Castellani LW, Kostem E, Furlotte N, Drake TA, Eskin E, Lusis AJ. A high-resolution Association mapping panel for the dissection of complex traits in mice. Genome Research. 2010;20:281–290. doi: 10.1101/gr.099234.109. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions

