The ribonuclease PNPase is a key regulator of biofilm formation in Listeria monocytogenes and affects invasion of host cells
- PMID: 37286543
- PMCID: PMC10247797
- DOI: 10.1038/s41522-023-00397-1
The ribonuclease PNPase is a key regulator of biofilm formation in Listeria monocytogenes and affects invasion of host cells
Abstract
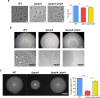
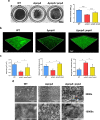
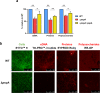
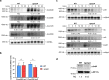
Biofilms provide an environment that protects microorganisms from external stresses such as nutrient deprivation, antibiotic treatments, and immune defences, thereby creating favorable conditions for bacterial survival and pathogenesis. Here we show that the RNA-binding protein and ribonuclease polynucleotide phosphorylase (PNPase) is a positive regulator of biofilm formation in the human pathogen Listeria monocytogenes, a major responsible for food contamination in food-processing environments. The PNPase mutant strain produces less biofilm biomass and exhibits an altered biofilm morphology that is more susceptible to antibiotic treatment. Through biochemical assays and microscopical analysis, we demonstrate that PNPase is a previously unrecognized regulator of the composition of the biofilm extracellular matrix, greatly affecting the levels of proteins, extracellular DNA, and sugars. Noteworthy, we have adapted the use of the fluorescent complex ruthenium red-phenanthroline for the detection of polysaccharides in Listeria biofilms. Transcriptomic analysis of wild-type and PNPase mutant biofilms reveals that PNPase impacts many regulatory pathways associated with biofilm formation, particularly by affecting the expression of genes involved in the metabolism of carbohydrates (e.g., lmo0096 and lmo0783, encoding PTS components), of amino acids (e.g., lmo1984 and lmo2006, encoding biosynthetic enzymes) and in the Agr quorum sensing-like system (lmo0048-49). Moreover, we show that PNPase affects mRNA levels of the master regulator of virulence PrfA and PrfA-regulated genes, and these results could help to explain the reduced bacterial internalization in human cells of the ΔpnpA mutant. Overall, this work demonstrates that PNPase is an important post-transcriptional regulator for virulence and adaptation to the biofilm lifestyle of Gram-positive bacteria and highlights the expanding role of ribonucleases as critical players in pathogenicity.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
The virulence regulator PrfA promotes biofilm formation by Listeria monocytogenes.J Bacteriol. 2010 Aug;192(15):3969-76. doi: 10.1128/JB.00179-10. Epub 2010 May 28. J Bacteriol. 2010. PMID: 20511507 Free PMC article.
-
PrfA led to reduced biofilm formation and contributed to altered gene expression patterns in biofilm-forming Listeria monocytogenes.Curr Microbiol. 2013 Sep;67(3):372-8. doi: 10.1007/s00284-013-0377-7. Epub 2013 May 8. Curr Microbiol. 2013. PMID: 23652633
-
AgrD-dependent quorum sensing affects biofilm formation, invasion, virulence and global gene expression profiles in Listeria monocytogenes.Mol Microbiol. 2009 Mar;71(5):1177-89. doi: 10.1111/j.1365-2958.2008.06589.x. Epub 2009 Jan 19. Mol Microbiol. 2009. PMID: 19154329
-
Molecular biology of surface colonization by Listeria monocytogenes: an additional facet of an opportunistic Gram-positive foodborne pathogen.Environ Microbiol. 2011 Apr;13(4):835-50. doi: 10.1111/j.1462-2920.2010.02378.x. Epub 2010 Nov 18. Environ Microbiol. 2011. PMID: 21087384 Review.
-
Listeria monocytogens - Amended understanding of its pathogenesis with a complete picture of its membrane vesicles, quorum sensing, biofilm and invasion.Microb Pathog. 2020 Dec;149:104575. doi: 10.1016/j.micpath.2020.104575. Epub 2020 Oct 20. Microb Pathog. 2020. PMID: 33091581 Review.
Cited by
-
Potential public health hazards related to consumption of poultry contaminated with antibiotic resistant Listeria monocytogenes in Egypt.BMC Microbiol. 2024 Jan 29;24(1):41. doi: 10.1186/s12866-024-03183-x. BMC Microbiol. 2024. PMID: 38287241 Free PMC article.
-
Integrative microbiome- and metatranscriptome-based analyses reveal diagnostic biomarkers for peri-implantitis.NPJ Biofilms Microbiomes. 2025 Aug 26;11(1):175. doi: 10.1038/s41522-025-00807-6. NPJ Biofilms Microbiomes. 2025. PMID: 40858628 Free PMC article.
References
-
- Shi X, Zhu X. Biofilm formation and food safety in food industries. Trends Food Sci. Technol. 2009;20:407–413. doi: 10.1016/j.tifs.2009.01.054. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

