Genome organization and genomics in Chlamydia: whole genome sequencing increases understanding of chlamydial virulence, evolution, and phylogeny
- PMID: 37287464
- PMCID: PMC10242142
- DOI: 10.3389/fcimb.2023.1178736
Genome organization and genomics in Chlamydia: whole genome sequencing increases understanding of chlamydial virulence, evolution, and phylogeny
Abstract
The genus Chlamydia contains important obligate intracellular bacterial pathogens to humans and animals, including C. trachomatis and C. pneumoniae. Since 1998, when the first Chlamydia genome was published, our understanding of how these microbes interact, evolved and adapted to different intracellular host environments has been transformed due to the expansion of chlamydial genomes. This review explores the current state of knowledge in Chlamydia genomics and how whole genome sequencing has revolutionised our understanding of Chlamydia virulence, evolution, and phylogeny over the past two and a half decades. This review will also highlight developments in multi-omics and other approaches that have complemented whole genome sequencing to advance knowledge of Chlamydia pathogenesis and future directions for chlamydial genomics.
Keywords: Chlamydia trachomatis; Chlamydiaceae; genome content; genomics; host tropism; next generation sequencing; tissue tropism; whole genome sequence.
Copyright © 2023 Luu, Kasimov, Phillips, Myers and Jelocnik.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

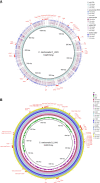




References
-
- Almeida F., Borges V., Ferreira R., Borrego M. J., Gomes J. P., Mota L. J. (2012). Polymorphisms in inc proteins and differential expression of inc genes among Chlamydia trachomatis strains correlate with invasiveness and tropism of lymphogranuloma venereum isolates. J. Bacteriol. 194 (23), 6574–6585. doi: 10.1128/jb.01428-12 - DOI - PMC - PubMed
-
- Andersson P., Harris S. R., Smith H. M. B. S., Hadfield J., O'Neill C., Cutcliffe L. T., et al. (2016). Chlamydia trachomatis from Australian aboriginal people with trachoma are polyphyletic composed of multiple distinctive lineages. Nat. Commun. 7, 10688–10688. doi: 10.1038/ncomms10688 - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

