Cell membrane sample preparation method of combined AFM and dSTORM analysis
- PMID: 37288003
- PMCID: PMC10185485
- DOI: 10.52601/bpr.2022.220004
Cell membrane sample preparation method of combined AFM and dSTORM analysis
Abstract
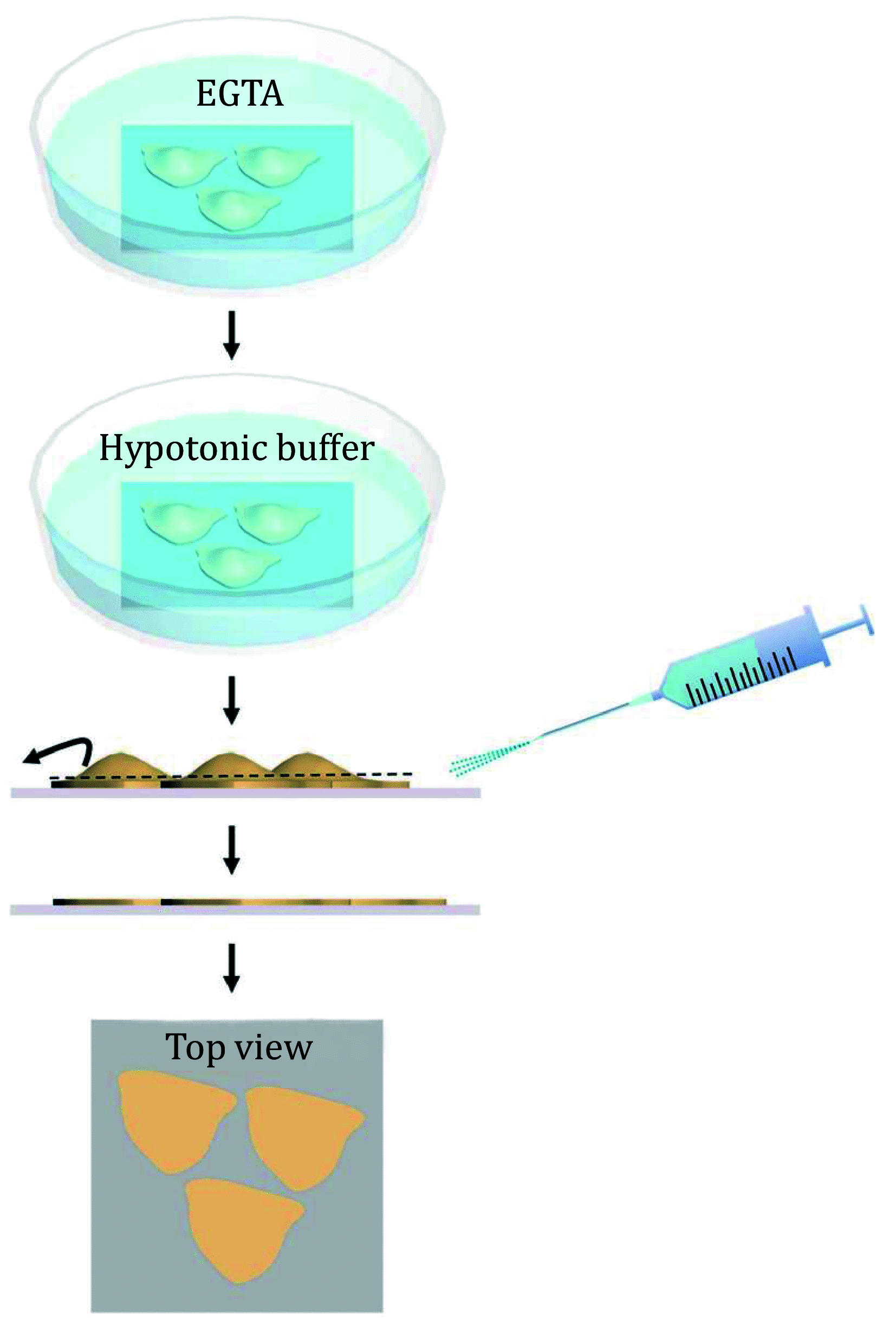
A major role of cell membranes is to provide an ideal environment for the constituent proteins to perform their biological functions. A deep understanding of the membrane proteins assembly process under physiological conditions is quite important to elucidate both the structure and the function of the cell membranes. Along these lines, in this work, a complete workflow of the cell membrane sample preparation and the correlated AFM and dSTORM imaging analysis methods are presented. A specially designed, angle-controlled sample preparation device was used to prepare the cell membrane samples. The correlated distributions of the specific membrane proteins with the topography of the cytoplasmic side of the cell membranes can be obtained by performing correlative AFM and dSTORM measurements. These methods are ideal for systematically studying the structure of the cell membranes. The proposed method of the sample characterization was not only limited to the measurement of the cell membrane but also can be applied for both biological tissue section analysis and detection.
Keywords: Atomic force microscopy (AFM); Cell membrane; Combined technique; Direct stochastic optical reconstruction microscopy (dSTORM); Super-resolution microscopy (SRM).
© The Author(s) 2022.
Conflict of interest statement
Mingjun Cai, Huili Wang, Guanfang Zhao, Hongru Li, Jing Gao and Hongda Wang declare that they have no conflict of interest.
Figures




Similar articles
-
High-Resolution Correlative Microscopy: Bridging the Gap between Single Molecule Localization Microscopy and Atomic Force Microscopy.Nano Lett. 2015 Aug 12;15(8):4896-904. doi: 10.1021/acs.nanolett.5b00572. Epub 2015 Jul 6. Nano Lett. 2015. PMID: 26121585
-
Correlative dual-color dSTORM/AFM reveals protein clusters at the cytoplasmic side of human bronchial epithelium membranes.Nanoscale. 2020 May 14;12(18):9950-9957. doi: 10.1039/c9nr10931e. Nanoscale. 2020. PMID: 32356532
-
Insight into the Different Channel Proteins of Human Red Blood Cell Membranes Revealed by Combined dSTORM and AFM Techniques.Anal Chem. 2021 Oct 26;93(42):14113-14120. doi: 10.1021/acs.analchem.1c02382. Epub 2021 Oct 17. Anal Chem. 2021. PMID: 34657412
-
How did correlative atomic force microscopy and super-resolution microscopy evolve in the quest for unravelling enigmas in biology?Nanoscale. 2021 Feb 4;13(4):2082-2099. doi: 10.1039/d0nr07203f. Nanoscale. 2021. PMID: 33346312 Review.
-
Atomic force microscopy of biological membranes.Biophys J. 2009 Jan;96(2):329-38. doi: 10.1016/j.bpj.2008.09.046. Biophys J. 2009. PMID: 19167286 Free PMC article. Review.
References
-
- Chacko JV, Zanacchi FC, Harke B, Lanzano L, Canale C, Diaspro A Insight into hybrid nanoscopy techniques: STED AFM & STORM AFM. Biophys J. 2014;106(2):396A–396A.
LinkOut - more resources
Full Text Sources
Miscellaneous
