This is a preprint.
A population-level computational histologic signature for invasive breast cancer prognosis
- PMID: 37293118
- PMCID: PMC10246230
- DOI: 10.21203/rs.3.rs-2947001/v1
A population-level computational histologic signature for invasive breast cancer prognosis
Update in
-
A population-level digital histologic biomarker for enhanced prognosis of invasive breast cancer.Nat Med. 2024 Jan;30(1):85-97. doi: 10.1038/s41591-023-02643-7. Epub 2023 Nov 27. Nat Med. 2024. PMID: 38012314
Abstract
Breast cancer is a heterogeneous disease with variable survival outcomes. Pathologists grade the microscopic appearance of breast tissue using the Nottingham criteria, which is qualitative and does not account for non-cancerous elements within the tumor microenvironment (TME). We present the Histomic Prognostic Signature (HiPS), a comprehensive, interpretable scoring of the survival risk incurred by breast TME morphology. HiPS uses deep learning to accurately map cellular and tissue structures in order to measure epithelial, stromal, immune, and spatial interaction features. It was developed using a population-level cohort from the Cancer Prevention Study (CPS)-II and validated using data from three independent cohorts, including the PLCO trial, CPS-3, and The Cancer Genome Atlas. HiPS consistently outperformed pathologists' performance in predicting survival outcomes, independent of TNM stage and pertinent variables. This was largely driven by stromal and immune features. In conclusion, HiPS is a robustly validated biomarker to support pathologists and improve prognosis.
Figures



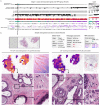


References
-
- American Cancer Society. Global Cancer Facts & Figures 4th Edition. Atlanta. (2018).
-
- Siegel R. L., Miller K. D., Wagle N. S. & Jemal A. Cancer statistics, 2023. CA Cancer J Clin 73, 17–48 (2023). - PubMed
-
- American Joint Commission on Cancer. AJCC Cancer Staging Manual 2017. (Springer International Publishing, 2017).
-
- Coughlin S. S. Social determinants of breast cancer risk, stage, and survival. Breast Cancer Res Treat 177, 537–548 (2019). - PubMed
-
- Li X. et al. Validation of the newly proposed American Joint Committee on Cancer (AJCC) breast cancer prognostic staging group and proposing a new staging system using the National Cancer Database. Breast Cancer Res Treat 171, 303–313 (2018). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

