A chromatin remodelling SWI/SNF subunit, Snr1, regulates neural stem cell determination and differentiation
- PMID: 37294080
- PMCID: PMC10323235
- DOI: 10.1242/dev.201484
A chromatin remodelling SWI/SNF subunit, Snr1, regulates neural stem cell determination and differentiation
Abstract
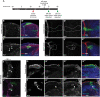
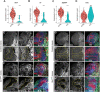
Coordinated spatio-temporal regulation of the determination and differentiation of neural stem cells is essential for brain development. Failure to integrate multiple factors leads to defective brain structures or tumour formation. Previous studies suggest changes of chromatin state are needed to direct neural stem cell differentiation, but the mechanisms are unclear. Analysis of Snr1, the Drosophila orthologue of SMARCB1, an ATP-dependent chromatin remodelling protein, identified a key role in regulating the transition of neuroepithelial cells into neural stem cells and subsequent differentiation of neural stem cells into the cells needed to build the brain. Loss of Snr1 in neuroepithelial cells leads to premature neural stem cell formation. Additionally, loss of Snr1 in neural stem cells results in inappropriate perdurance of neural stem cells into adulthood. Snr1 reduction in neuroepithelial or neural stem cells leads to the differential expression of target genes. We find that Snr1 is associated with the actively transcribed chromatin region of these target genes. Thus, Snr1 likely regulates the chromatin state in neuroepithelial cells and maintains chromatin state in neural stem cells for proper brain development.
Keywords: Drosophila; Differentiation; Neural stem cell; Neuroblast; Neuroepithelial cells; Optic lobe; SMARCB1; SWI/SNF complex; Snr1.
© 2023. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures






References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

