Characterization of the Inflammatory Response Evoked by Bacterial Membrane Vesicles in Intestinal Cells Reveals an RIPK2-Dependent Activation by Enterotoxigenic Escherichia coli Vesicles
- PMID: 37306596
- PMCID: PMC10433812
- DOI: 10.1128/spectrum.01115-23
Characterization of the Inflammatory Response Evoked by Bacterial Membrane Vesicles in Intestinal Cells Reveals an RIPK2-Dependent Activation by Enterotoxigenic Escherichia coli Vesicles
Abstract
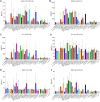
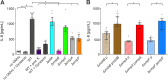
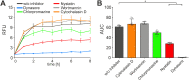
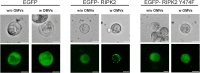
Although the immunomodulatory potency of bacterial membrane vesicles (MVs) is widely acknowledged, their interactions with host cells and the underlying signaling pathways have not been well studied. Herein, we provide a comparative analysis of the proinflammatory cytokine profile secreted by human intestinal epithelial cells exposed to MVs derived from 32 gut bacteria. In general, outer membrane vesicles (OMVs) from Gram-negative bacteria induced a stronger proinflammatory response than MVs from Gram-positive bacteria. However, the quality and quantity of cytokine induction varied between MVs from different species, highlighting their unique immunomodulatory properties. OMVs from enterotoxigenic Escherichia coli (ETEC) were among those showing the strongest proinflammatory potency. In depth analyses revealed that the immunomodulatory activity of ETEC OMVs relies on a so far unprecedented two-step mechanism, including their internalization into host cells followed by intracellular recognition. First, OMVs are efficiently taken up by intestinal epithelial cells, which mainly depends on caveolin-mediated endocytosis as well as the presence of the outer membrane porins OmpA and OmpF on the MVs. Second, lipopolysaccharide (LPS) delivered by OMVs is intracellularly recognized by novel caspase- and RIPK2-dependent pathways. This recognition likely occurs via detection of the lipid A moiety as ETEC OMVs with underacylated LPS exhibited reduced proinflammatory potency but similar uptake dynamics compared to OMVs derived from wild-type (WT) ETEC. Intracellular recognition of ETEC OMVs in intestinal epithelial cells is pivotal for the proinflammatory response as inhibition of OMV uptake also abolished cytokine induction. The study signifies the importance of OMV internalization by host cells to exercise their immunomodulatory activities. IMPORTANCE The release of membrane vesicles from the bacterial cell surface is highly conserved among most bacterial species, including outer membrane vesicles (OMVs) from Gram-negative bacteria as well as vesicles liberated from the cytoplasmic membrane of Gram-positive bacteria. It is becoming increasingly evident that these multifactorial spheres, carrying membranous, periplasmic, and even cytosolic content, contribute to intra- and interspecies communication. In particular, gut microbiota and the host engage in a myriad of immunogenic and metabolic interactions. This study highlights the individual immunomodulatory activities of bacterial membrane vesicles from different enteric species and provides new mechanistic insights into the recognition of ETEC OMVs by human intestinal epithelial cells.
Keywords: ETEC; HT-29; IL-8; LPS; OMV; RIPK2; bacterial membrane vesicles; caspase; cytokine; cytokines; inflammatory response; interleukins; intestinal epithelial cell; lipopolysaccharide; outer membrane vesicles; porin; porins.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


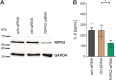
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

