Trophoblast PR-SET7 dysfunction induces viral mimicry response and necroptosis associated with recurrent miscarriage
- PMID: 37307441
- PMCID: PMC10288560
- DOI: 10.1073/pnas.2216206120
Trophoblast PR-SET7 dysfunction induces viral mimicry response and necroptosis associated with recurrent miscarriage
Abstract
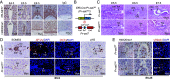
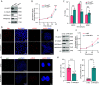
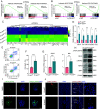
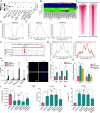
Recurrent miscarriage (RM) is a distressing pregnancy complication. While the etiology of RM remains unclear, growing evidence has indicated the relevance of trophoblast impairment to the pathogenesis of RM. PR-SET7 is the sole enzyme catalyzing monomethylation of H4K20 (H4K20me1) and has been implicated in many pathophysiological processes. However, how PR-SET7 functions in trophoblasts and its relevance to RM remain unknown. Here, we found that trophoblast-specific loss of Pr-set7 in mice led to defective trophoblasts, resulting in early embryonic loss. Mechanistic analysis revealed that PR-SET7 deficiency in trophoblasts derepressed endogenous retroviruses (ERVs), leading to double-stranded RNA stress and subsequent viral mimicry, which drove overwhelming interferon response and necroptosis. Further examination discovered that H4K20me1 and H4K20me3 mediated the inhibition of cell-intrinsic expression of ERVs. Importantly, dysregulation of PR-SET7 expression and the corresponding aberrant epigenetic modifications were observed in the placentas of RM. Collectively, our results demonstrate that PR-SET7 acts as an epigenetic transcriptional modulator essential for repressing ERVs in trophoblasts, ensuring normal pregnancy and fetal survival, which sheds new light on potential epigenetic causes contributing to RM.
Keywords: ERVs; H4K20me1/3; PR-SET7; recurrent miscarriage; viral mimicry.
Conflict of interest statement
The authors declare no competing interest.
Figures


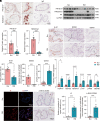

References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

